Selecting Molecular Descriptors for QSAR: A Strategic Guide from Foundations to AI-Enhanced Validation
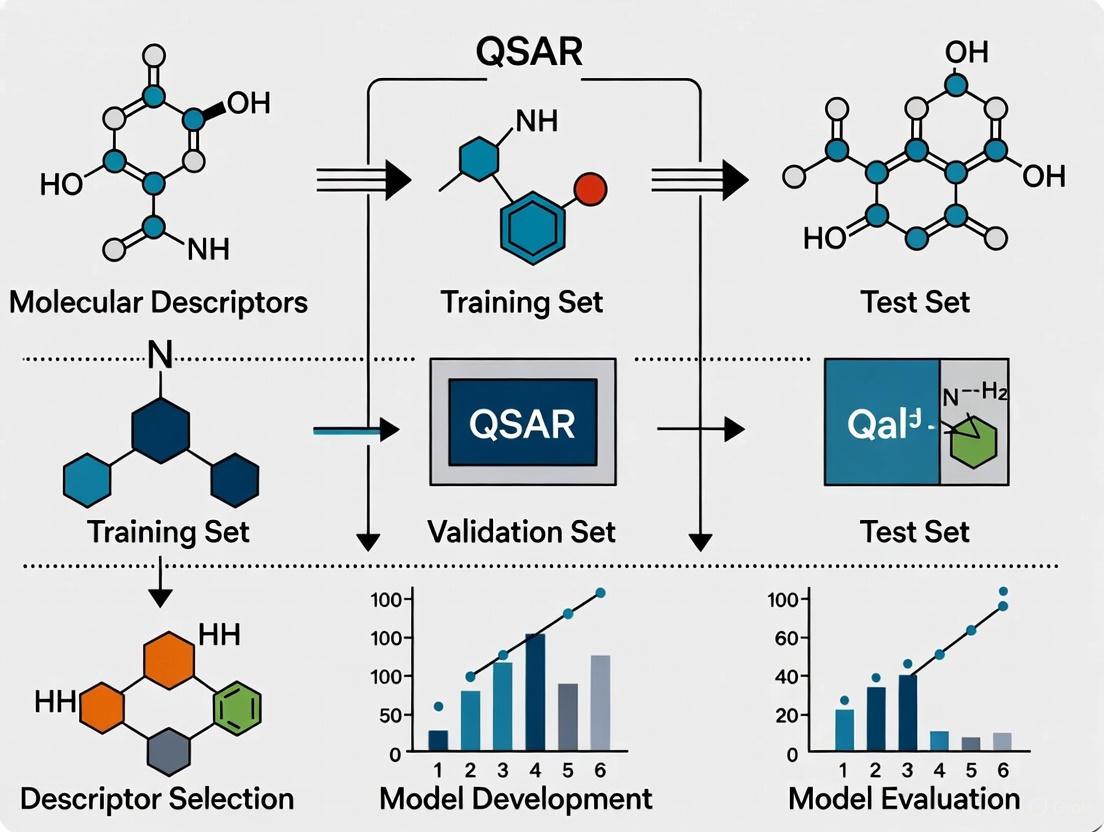
This article provides a comprehensive guide for researchers and drug development professionals on the critical process of selecting molecular descriptors for Quantitative Structure-Activity Relationship (QSAR) modeling.
Selecting Molecular Descriptors for QSAR: A Strategic Guide from Foundations to AI-Enhanced Validation
Abstract
This article provides a comprehensive guide for researchers and drug development professionals on the critical process of selecting molecular descriptors for Quantitative Structure-Activity Relationship (QSAR) modeling. It covers the foundational principles of molecular descriptors, from 1D physicochemical properties to 4D conformational ensembles and AI-generated deep descriptors. The piece delves into methodological strategies, including variable selection techniques and the impact of descriptor choice on model interpretability. It further addresses common troubleshooting scenarios, such as managing high-dimensional data and defining the model's applicability domain. Finally, the article synthesizes modern validation paradigms, comparing classical and machine learning approaches, and emphasizes the necessity of rigorous external validation and adherence to OECD principles for developing robust, predictive QSAR models in drug discovery.
The Building Blocks of QSAR: Understanding Molecular Descriptor Types and Their Fundamental Roles
Frequently Asked Questions (FAQs)
Q1: What is the fundamental difference between a 1D and a 4D molecular descriptor? The core difference lies in the complexity of the molecular representation they capture. A 1D descriptor typically represents global, whole-molecule properties that do not require structural or connectivity information, such as molecular weight or atom counts [1] [2]. In contrast, a 4D descriptor incorporates the dimension of time and interaction fields, often derived from molecular dynamics simulations or the placement of a molecule within a 3D grid to probe its interactions with a receptor site, providing information on specific, conformation-dependent interactions [1] [2].
Q2: My QSAR model is overfitting. How can my choice of descriptors contribute to this, and how can I address it? Overfitting often occurs when the number of descriptors is too large relative to the number of compounds in your dataset, or when descriptors are highly correlated [3]. To address this:
- Apply Feature Selection: Use techniques like genetic algorithms (wrapper methods) or LASSO regression (embedded methods) to identify and retain only the most relevant descriptors [3].
- Start Simpler: Consider beginning your modeling process with simpler, more interpretable 1D or 2D descriptors, which can be just as predictive as 3D/4D descriptors for many endpoints and are less computationally intensive [2].
- Validate Rigorously: Always use robust validation methods, such as k-fold cross-validation and an external test set, to ensure your model's performance is genuine [4] [3].
Q3: I've identified an important "bulk" property descriptor like molecular weight in my model. Should I use this to guide chemical modifications? Not necessarily in isolation. Recent research highlights that high-dimensional descriptor spaces are often confounded, meaning a "bulk" property may be a proxy for a true, specific pharmacophore [5]. Before guiding synthesis, it is crucial to perform deconfounding analysis to determine if the descriptor has a causal link to the activity, or merely a correlational one. Advanced statistical frameworks, such as Double Machine Learning (DML), can help distinguish true causal features from spurious ones [5].
Q4: How do I know which level of descriptor (1D-4D) to start with for my QSAR project? A hierarchical approach is often most efficient [6]:
- Start with 1D/2D Descriptors: These are fast to compute and often provide a strong baseline model. If performance is satisfactory, this can save significant time and resources [2].
- Progress to 3D Descriptors: If the biological endpoint is known to be highly dependent on stereochemistry or 3D shape (e.g., receptor binding), move to 3D descriptors. Ensure you have a reliable method for conformational sampling [2].
- Reserve 4D for Complex Problems: Use 4D descriptors for the most challenging cases where explicit modeling of ligand-receptor interactions or dynamics is necessary [6].
Q5: What are the minimal criteria for a molecular descriptor to be considered well-defined and useful for QSAR? A robust molecular descriptor should meet several key criteria [1]:
- Invariance: Its value must be invariant to molecular manipulations that don't change the underlying structure, such as atom numbering, rotation, or translation.
- Unambiguous Algorithm: It must be defined by a clear and unambiguous mathematical procedure.
- Good Correlation: It should correlate with at least one experimental property.
- Low Degeneracy: It should, ideally, have a low probability of producing the same value for different molecules.
- Structural Interpretation: The descriptor should have a meaningful chemical or structural interpretation to provide insights for chemists [7].
Troubleshooting Common Experimental Issues
Problem: Poor Predictive Performance on External Test Set Your model performs well on training data but poorly on new, unseen compounds.
| Possible Cause | Diagnostic Steps | Solution |
|---|---|---|
| Incorrect Applicability Domain | Check if the new compounds are structurally dissimilar to your training set. | Define the applicability domain of your model. Use similarity metrics to ensure new compounds fall within the chemical space the model was trained on [3]. |
| Data Quality Issues | Re-inspect the original experimental data for the training set. Look for errors, outliers, or inconsistent measurement conditions. | Perform rigorous data cleaning and curation: standardize structures, remove duplicates, and handle missing values appropriately [3]. |
| Overfitting | Compare performance metrics between the training set and cross-validation. A large gap indicates overfitting. | Apply feature selection to reduce the number of descriptors and simplify the model. Use regularization techniques [3]. |
Problem: Model Lacks Chemical Interpretability The model is predictive, but you cannot extract meaningful chemical insights to guide design.
| Possible Cause | Diagnostic Steps | Solution |
|---|---|---|
| Use of "Black Box" Models/Descriptors | Evaluate the model's inherent interpretability. Models like Random Forest can provide feature importance. | Use descriptor types that are chemically intuitive. Implement model interpretation techniques like the Gini index for Random Forest to identify which structural features (e.g., aromatic moieties, specific atoms) are most influential [4] [7]. |
| High Correlation Among Descriptors | Calculate the correlation matrix of your descriptors. | Apply a descriptor whitening technique or select a subset of uncorrelated descriptors to isolate the individual effect of each feature. Consider causal inference methods to deconfound descriptors [5]. |
The Scientist's Toolkit: Essential Software for Descriptor Calculation
The following table lists key software tools for computing molecular descriptors, along with their capabilities and key characteristics.
| Software Name | 0D/1D | 2D Fingerprints | 3D/4D Descriptors | Key Characteristics | License |
|---|---|---|---|---|---|
| alvaDesc [1] | Yes | Yes | Yes | Comprehensive descriptor calculation; available for Windows, Linux, macOS; updated in 2025. | Proprietary, Commercial |
| Dragon [1] | Yes | Yes | Yes | Historically a industry standard; now discontinued. | Proprietary, Commercial |
| Mordred [1] | Yes | No | Yes | Based on RDKit; open-source; a community-maintained fork is available. | Free, Open Source |
| PaDEL-Descriptor [1] [3] | Yes | Yes | Yes | Based on the Chemistry Development Kit (CDK); discontinued but widely used. | Free |
| RDKit [1] [3] | Yes | Yes | Yes | Versatile cheminformatics toolkit; includes descriptor calculation; actively updated (2024). | Free, Open Source |
| scikit-fingerprints [1] | Yes | Yes | Yes | A Python library specifically for calculating molecular fingerprints; updated in 2025. | Free, Open Source |
Standard Experimental Protocol: A Hierarchical Workflow for Descriptor Selection and Model Building
This protocol outlines a systematic, hierarchical approach for selecting molecular descriptors and developing a validated QSAR model, based on established best practices [3] [6].
Objective: To build a robust and interpretable QSAR model by sequentially progressing through levels of molecular complexity, using the information from each step to inform the next.
Materials:
- Dataset: A curated set of chemical structures (e.g., as SMILES strings) and their corresponding biological activity values (e.g., IC50, pIC50).
- Software: A descriptor calculation tool (see Toolkit table above), a data analysis environment (e.g., Python with scikit-learn, R), and a molecular modeling platform if 3D/4D descriptors are required.
Methodology:
Step 1: Data Curation and Preparation
- Collect and Clean: Compile the dataset from reliable sources (e.g., ChEMBL, PubChem). Standardize chemical structures: remove salts, normalize tautomers, and define stereochemistry clearly [3].
- Handle Activity Data: Convert all activity values to a consistent scale (e.g., log-transform IC50 to pIC50).
- Split Data: Divide the dataset into a training set (~80%) and a hold-out test set (~20%). The test set must be kept completely separate until the final model validation [3].
Step 2: Hierarchical Descriptor Calculation and Modeling The essence of this hierarchic system is that the QSAR problem is solved sequentially. At each subsequent stage, the problem is not solved from scratch, but the information obtained from the previous step is used [6].
1D/2D Model:
- Calculate Descriptors: Compute a large set of 1D (constitutional) and 2D (topological) descriptors using software like RDKit or Mordred.
- Feature Selection: Apply a feature selection method (e.g., genetic algorithm, LASSO) to the training set to reduce dimensionality and avoid overfitting.
- Model Building & Validation: Train a model (e.g., Random Forest, PLS) using the selected descriptors. Evaluate performance using 5-fold cross-validation on the training set. Record key metrics (e.g., R², Q², MCC).
3D Model:
- Generate 3D Conformers: For each molecule, generate a low-energy 3D conformation.
- Calculate 3D Descriptors: Compute 3D descriptors (e.g., WHIM, GETAWAY, quantum-chemical descriptors).
- Feature Selection & Modeling: Perform feature selection on the 3D descriptor pool. Build and validate a new model using the same protocol as in Step 2.1.
- Compare and Learn: Compare the performance of the 3D model with the 1D/2D model. If performance does not improve significantly, the simpler model may be sufficient. Analyze the important 3D descriptors for structural insights [2].
4D Model (If Required):
- Define Interaction Probes: Place the molecules in a 3D grid and use probes (e.g., water, methyl group) to calculate interaction energy fields (as in 4D descriptors or CoMFA) [2].
- Calculate 4D Descriptors: Derive descriptors from these interaction fields or from molecular dynamics trajectories.
- Build Final Model: Build and validate a model using these 4D descriptors. This step is typically reserved for cases where 3D shape and interaction specifics are critical and simpler models are inadequate.
Step 3: Final Model Validation and Reporting
- External Validation: Apply the final chosen model (whether it be from 1D/2D, 3D, or 4D) to the hold-out test set that was set aside in Step 1. This provides an unbiased estimate of the model's predictive power [3].
- Define Applicability Domain: Characterize the chemical space of the training set to establish the range of structures for which the model can make reliable predictions [3].
- Report Results: Document the model's algorithm, key descriptors, performance statistics, and applicability domain according to regulatory standards like the OECD QSAR Model Reporting Format (QMRF) where applicable [8] [9].
Hierarchical Descriptor Selection Workflow
Data Presentation: Key Performance Metrics from a Modern QSAR Study
The following table summarizes the robust performance of a Random Forest QSAR model using SubstructureCount fingerprints, developed to predict the activity of Plasmodium falciparum dihydroorotate dehydrogenase (PfDHODH) inhibitors for anti-malarial drug discovery [4]. The model was validated on balanced data using an oversampling technique.
| Model Stage | Matthews Correlation Coefficient (MCC) | Accuracy | Sensitivity (Recall) | Specificity |
|---|---|---|---|---|
| Training Set | 0.97 | > 80% | > 80% | > 80% |
| Cross-Validation | 0.78 | > 80% | > 80% | > 80% |
| External Test Set | 0.76 | > 80% | > 80% | > 80% |
Interpretation: The high MCC values across all stages, particularly the strong external test set MCC of 0.76, indicate a model with excellent predictive power and robustness, minimizing false positives and false negatives. The high sensitivity and specificity confirm its balanced ability to identify both active and inactive compounds [4].
Core Concepts: The Dimensionality of Molecular Descriptors
Hierarchy of Molecular Descriptor Dimensions
Frequently Asked Questions (FAQs) on Descriptor Selection and Application
Q1: What is the fundamental difference between classical Hansch analysis and modern descriptor-based QSAR?
A1: Classical Hansch Analysis is a Linear Free Energy Relationship (LFER) approach that uses a limited set of interpretable physicochemical parameters—namely hydrophobicity (Log P), electronic effects (Hammett σ constants), and steric effects (Taft Es constants)—to correlate structure with biological activity via a linear equation [10]. In contrast, modern QSAR utilizes high-dimensional molecular descriptors (often hundreds or thousands) and advanced machine learning (ML) algorithms. The key challenge with modern approaches is that standard ML models can be misled by high correlations between these descriptors, incorrectly identifying proxy "bulk" properties (e.g., molecular weight) as important, instead of the true causal pharmacophoric features [5].
Q2: My QSAR model has good internal validation statistics but fails in external prediction. What could be the cause?
A2: This is a common issue often rooted in experimental errors within the training data and overfitting. Studies show that even a small ratio of experimental errors in the modeling set can significantly deteriorate external prediction performance [11]. While consensus predictions can help identify compounds with potential experimental errors, simply removing compounds with large cross-validation errors does not reliably improve external predictivity and may lead to overfitting [11]. Furthermore, models trained on confounded correlations rather than true causal effects are likely to fail when applied to new chemical spaces [5].
Q3: How can I identify and mitigate the effect of experimental errors in my dataset?
A3: You can use the QSAR modeling process itself to help prioritize potential outliers. The methodology is as follows [11]:
- Develop a QSAR model and perform a cross-validation (e.g., fivefold).
- Sort all compounds in the dataset by the magnitude of their prediction errors from the cross-validation.
- Analyze the top compounds with the largest errors; these are likely to contain experimental inaccuracies. Research indicates that this method can successfully prioritize compounds with simulated experimental errors, with performance quantified by ROC enrichment factors [11]. Table: Experimental Error Prioritization Performance
| Dataset Type | Top 1% Enrichment | Top 20% Enrichment | Notes |
|---|---|---|---|
| Categorical (MDR1) | 12.9x | 4.7x | Compared to random selection |
| Continuous (LD50) | 4.2x - 5.3x | 2.3x | Varies by error simulation strategy |
Q4: What are "causal descriptors," and how can they be identified?
A4: Causal descriptors are molecular features that have a statistically significant and unconfounded causal effect on the biological activity, rather than just a correlational link. A framework using Double/Debiased Machine Learning (DML) has been proposed to identify them [5]. The experimental protocol involves:
- DML Estimation: Use DML to estimate the causal effect of each individual molecular descriptor on the activity, while treating all other p-1 descriptors as potential confounders.
- Hypothesis Testing: Apply statistical testing (e.g., the Benjamini-Hochberg procedure) to the p causal estimates to control the False Discovery Rate (FDR). This framework has been shown in validation studies to successfully reject spurious, confounded descriptors and correctly identify the true causal features [5].
Troubleshooting Guides for Common Technical Issues
Issue 1: Poor Model Interpretability and Spurious Correlations
Problem: The model highlights "bulk" properties like molecular weight as key drivers, which are likely proxies, not true mechanistic features.
Solution: Implement a causal inference framework.
- Apply Deconfounding Techniques: Use the DML and FDR control framework to deconfound the descriptor space [5].
- Workflow: The diagram below outlines the process for identifying causal descriptors.
Issue 2: Handling Structural and Experimental Data Errors
Problem: Underlying data quality issues, such as structural misrepresentation or experimental variability, lead to poor and unreliable models.
Solution: Establish a rigorous data curation and validation protocol.
- Chemical Structure Curation: Follow a standardized workflow to remove and correct structural errors, which is a major source of model inaccuracy [11].
- Experimental Error Identification: Use the cross-validation prioritization method described in FAQ A3 to flag potential outliers for expert review [11].
- Leverage Consensus: Employ consensus predictions from multiple models, as they are more robust in identifying compounds with potential experimental errors [11].
Issue 3: Integrating Read-Across with QSAR Modeling
Problem: Simple read-across predictions are subjective and hard to quantify, while QSAR models may lack sufficient data.
Solution: Combine the strengths of both approaches using novel methodologies.
- Use RASAR Frameworks: Implement Read-Across Structure-Activity Relationship (RASAR) models. These use similarity-based descriptors derived from read-across, combined with traditional molecular descriptors, to build more predictive ML models [12].
- Adopt Advanced Platforms: Utilize platforms like OrbiTox, which integrate chemistry-based similarity searching, molecular descriptors, and QSAR models into a unified read-across workflow [13]. Frameworks like Generalized Read-Across (GenRA) also provide a more quantitative and automated approach [12].
- Workflow: The following diagram illustrates the integrated RASAR approach.
Table: Key Resources for Evolving QSAR Practices
| Tool/Resource | Category | Function & Explanation |
|---|---|---|
| Hansch Equation | Foundational Model | The original framework relating biological activity (log 1/C) to hydrophobicity (log P), electronic (σ), and steric (Es) parameters [10]. |
| Double Machine Learning (DML) | Statistical Method | A causal inference method used to deconfound molecular descriptors and estimate true causal effects on activity [5]. |
| Benjamini-Hochberg Procedure | Statistical Method | A hypothesis testing procedure used to control the False Discovery Rate (FDR) when testing hundreds of molecular descriptors simultaneously [5]. |
| Read-Across Structure-Activity Relationship (RASAR) | Modeling Approach | A hybrid technique that uses similarity descriptors from read-across to build more predictive QSAR-like models [12]. |
| OrbiTox Platform | Software Platform | A read-across platform featuring similarity searching, Saagar molecular descriptors, and built-in QSAR models for regulatory submissions [13]. |
| OECD QSAR Toolbox | Software Platform | A widely used software for profiling chemicals, filling data gaps via read-across, and grouping chemicals into categories [8]. |
| Consensus Modeling | Modeling Strategy | Averaging predictions from multiple individual QSAR models to improve robustness and identify potential experimental errors [11]. |
Fundamental Concepts FAQ
What are lipophilicity (Log P), electronic, and steric effects and why are they crucial for QSAR?
Lipophilicity, commonly measured as the partition coefficient Log P, quantifies how a compound distributes itself between a lipophilic phase (like octanol) and an aqueous phase (like water). It is a key determinant in a drug's absorption, distribution, membrane permeability, and overall pharmacokinetics [14] [15]. According to Lipinski's "rule of five," an orally active drug candidate should typically have a Log P value of less than 5 [14]. For ionizable compounds, the distribution coefficient Log D (which accounts for all ionized and unionized species) is used instead, as it provides a more accurate picture at physiological pH values [14] [15].
Electronic effects describe how the electron distribution within a molecule influences its interactions. This includes the influence of lone-pair electrons, atomic charges, and molecular orbital energies (like HOMO and LUMO), which affect a molecule's polarity, polarizability, and its ability to form hydrogen bonds [16] [17]. These factors are critical for understanding binding interactions with a biological target.
Steric effects relate to the spatial arrangement and bulkiness of atoms within a molecule, which can physically impede interactions with a biological target. Steric parameters help quantify molecular volume and shape, which are vital for understanding how a drug fits into its binding site [16] [18].
In QSAR, these properties are translated into molecular descriptors. They form the foundation of models that connect a molecule's physical structure to its biological activity, enabling the prediction and optimization of new drug candidates [14] [16].
What is the difference between Log P and Log D?
| Property | Definition | Best Used For |
|---|---|---|
| Log P | The logarithm of the partition coefficient for the uncharged, neutral form of a molecule between octanol and water [14]. | Non-ionic compounds; a pure measure of intrinsic lipophilicity. |
| Log D | The logarithm of the apparent distribution coefficient, which accounts for all forms of the compound (both ionized and unionized) in the two phases at a specific pH [14] [15]. | Ionizable compounds; provides a more relevant measure of lipophilicity at a given physiological pH (e.g., 7.4 for blood). |
The relationship between Log D and Log P for ionizable compounds is given by: Log D = Log P - log(1 + 10^(pH-pKa)) for acids, and with a corresponding adjustment for bases [14]. This highlights that Log D is pH-dependent, making it essential for modeling activity across different physiological environments like the stomach (pH ~2) or intestine (pH 5-6.8) [14].
Troubleshooting Guides
How do I select the right molecular descriptors for my QSAR model?
Problem: Model shows poor predictive power, potentially due to inappropriate descriptor selection.
Solution: Follow this systematic workflow to choose descriptors based on your molecular property and available resources.
Why does my calculated Log P differ from experimental values, and how can I improve accuracy?
Problem: Significant discrepancy between computational Log P predictions and experimental shake-flask results.
Solution:
Identify the Source of Error:
- Ionizable Compounds: Ensure you are calculating Log D, not Log P, if your compound ionizes within the physiological pH range. This is a common oversight [14] [15].
- Molecular Complexity: Fragment-based calculation methods can be inaccurate for complex molecules with intramolecular interactions (e.g., hydrogen bonding) that are not fully captured by simple group contributions [15].
- Experimental Error: The experimental data in your training set may itself contain errors, which can skew your model [11].
Troubleshooting Steps:
| Step | Action | Rationale |
|---|---|---|
| 1 | Verify the ionization state (pKa) of your compound and calculate Log D at the relevant pH. | Corrects for the most common error in lipophilicity assessment for ionizable drugs [14]. |
| 2 | Use a consensus prediction by averaging results from multiple computational methods (fragment-based, whole-molecule, etc.). | Mitigates the inherent limitations and biases of any single calculation method [11]. |
| 3 | For critical compounds, validate computational predictions with a high-throughput experimental measure like HPLC retention time comparison. | Provides an experimental anchor point and helps identify outliers in computational predictions [15]. |
| 4 | Cross-validate your QSAR model and check if the compound is within the model's Applicability Domain (AD). | Flags predictions for molecules that are too structurally dissimilar from the training set, which are likely to be unreliable [11]. |
How can I computationally quantify electronic and steric effects for my QSAR study?
Problem: Need robust, calculable descriptors for electronic and steric properties.
Solution: Utilize the following descriptors, selectable based on your computational resources and need for accuracy.
Table: Computational Descriptors for Electronic and Steric Effects
| Effect Type | Descriptor | Description | Calculation Method & Notes |
|---|---|---|---|
| Electronic | HOMO/LUMO Energies | Energy of the Highest Occupied and Lowest Unoccupied Molecular Orbitals. Indicates nucleophilicity/electrophilicity [17]. | Quantum Chemical Calculation (DFT, Semi-empirical). A fundamental QM descriptor. |
| Atomic Partial Charges | The calculated electron density on individual atoms. | Semi-empirical or DFT. Can be used in regression equations for Log P [14]. | |
| Lone-Pair Electron Index (LEI) | A topological index that quantifies the electrostatic effect of heteroatoms' lone-pair electrons [16]. | Topological/Fragment-based. Simple to calculate and highly effective in QSAR models [16]. | |
| Dipole Moment | Measure of the overall molecular polarity. | Quantum Chemical Calculation. Influenced by both molecular symmetry and atomic charges. | |
| Steric | Molecular Volume Index (MVI) | A topological index based on van der Waals volumes of atoms [16]. | Topological. Easy to compute from molecular structure. |
| Taft's Steric Parameter (Eₛ) | A classic parameter defining the bulk of a substituent [16] [18]. | Empirical/Fragment-based. Derived from experimental kinetics; available from lookup tables. | |
| van der Waals Volume | The 3D volume occupied by the molecule. | Quantum Chemical or Molecular Mechanics. Provides a direct 3D measure of molecular bulk. |
Experimental Protocols & Methodologies
Protocol: Predicting Log P via Direct Solvation Free Energy Calculation
This method uses quantum chemical calculations combined with continuum solvation models to predict Log P based on first principles [14].
Principle: Log P is calculated from the transfer free energy (ΔGtransfer) of a molecule from water to octanol, using the formula: log P = -ΔGtransfer / (RT ln 10), where ΔGtransfer = ΔGsolvation(octanol) - ΔG_solvation(water) [14].
Workflow:
- Geometry Optimization: Perform a full geometry optimization of the molecule in the gas phase using a quantum chemical method (e.g., Density Functional Theory with the B3LYP functional and 6-31G* basis set) [19] [17].
- Solvation Free Energy Calculation: Using the optimized geometry, calculate the solvation free energy (ΔGsolvation) in two separate single-point energy calculations:
- ΔGsolvation(water): In a continuum solvation model representing water (e.g., IEF-PCM or SMD).
- ΔG_solvation(octanol): In a continuum solvation model representing octanol.
- Compute Log P: Calculate ΔG_transfer and apply the formula above to obtain the Log P value.
Troubleshooting Tips:
- High Computational Cost: For large datasets, use faster semi-empirical quantum methods (e.g., PM6/MOPAC) as a trade-off between speed and accuracy [17].
- Inaccurate for Ions: This direct method is best suited for neutral molecules. For ions, use the Log D relationship [14].
Protocol: Calculating Quantum Chemical Electronic Descriptors
This protocol outlines the steps to compute key electronic descriptors like HOMO/LUMO energies and polarizability [17].
Software: Use quantum chemical software packages like Gaussian, GAMESS, or MOPAC, often with a graphical interface like MOLDEN.
Step-by-Step Guide (e.g., for HOMO Energy):
- Build the Molecule: Construct a 3D model using a molecular builder.
- Submit Geometry Optimization: Run a geometry optimization job to find the molecule's most stable structure. A common method is DFT with the B3LYP functional and the 6-31G* basis set [19].
- Analyze Output:
- Once optimized, the output file contains the orbital energies.
- Open the output in a program like MOLDEN to visualize the HOMO and LUMO orbitals to confirm their character.
- The HOMO and LUMO energies are directly listed in the output file [17].
Step-by-Step Guide for Polarizability:
- Start with Optimized Geometry: Use the geometry from Step 2 above.
- Submit a Single-Point Calculation: Run a calculation with the
POLARkeyword (in MOPAC) or a similar function to request polarizability calculation. - Extract Result: The polarizability volume (in ų) is reported in the output file [17].
The Scientist's Toolkit: Research Reagent Solutions
Table: Essential Computational and Experimental Resources
| Item Name | Function in Research | Application Context |
|---|---|---|
| clogP Software | Fragment-based calculation of Log P for high-throughput virtual screening [14]. | Rapid prediction of lipophilicity in early-stage drug discovery. |
| Continuum Solvation Model (e.g., IEF-PCM) | A computational model that treats the solvent as a continuous dielectric to calculate solvation free energies [14]. | Used for direct, QM-based Log P prediction and solvation energy calculations. |
| Quantum Chemical Software (Gaussian, GAMESS) | Performs ab initio or DFT calculations to compute electronic descriptors (HOMO/LUMO, charges, polarizability) [17]. | Generating highly accurate electronic structure descriptors for QSAR. |
| Semi-empirical Software (MOPAC) | Uses parameterized quantum methods for faster calculation of properties for large molecules [17]. | A balance of speed and accuracy for larger datasets or molecules. |
| n-Octanol/Water System | The experimental gold-standard system for measuring Log P via the shake-flask method [15]. | Generating experimental lipophilicity data for validation. |
| Immobilized Artificial Membrane (IAM) | Chromatographic surface that mimics a cell membrane to measure drug-membrane partitioning [15]. | Provides a more biologically relevant measure of lipophilicity than octanol/water. |
FAQs: Core Definitions and Selection
1. What are the fundamental differences between topological, quantum chemical, and 3D surface descriptors?
Topological, quantum chemical, and 3D surface descriptors encode different aspects of molecular structure, making them suitable for various applications in Quantitative Structure-Activity Relationship (QSAR) modeling. The table below summarizes their core characteristics.
Table 1: Fundamental Comparison of Molecular Descriptor Types
| Descriptor Type | Definition & Basis | Key Examples | Primary Applications in QSAR |
|---|---|---|---|
| Topological Descriptors | 2D numerical indices encoding molecular connectivity and atomic arrangement from the molecular graph. [20] | Wiener index, Zagreb indices, Connectivity index ( [21]) | Modeling molecular size, shape, branching; high-throughput virtual screening of large databases. [20] [21] |
| Quantum Chemical Descriptors | Descriptors derived from quantum mechanical calculations, representing electronic structure and energetic properties. [22] | HOMO/LUMO energies, Hardness (η), Electrostatic Potential (ESP), Polarizability (α) [22] | Predicting chemical reactivity, reaction mechanisms, and interactions involving electron transfer. [22] [23] |
| 3D Surface Descriptors | Descriptors based on the molecule's 3D structure, representing steric and electrostatic fields around it. [24] | Comparative Molecular Field Analysis (CoMFA), Comparative Molecular Similarity Indices Analysis (CoMSIA) fields [24] | Understanding steric and electrostatic requirements for ligand-receptor binding; lead optimization. [24] |
2. When should I prioritize quantum chemical descriptors over topological descriptors?
Prioritize quantum chemical descriptors when your research involves predicting or interpreting phenomena directly related to a molecule's electronic structure, such as [22] [23]:
- Chemical reactivity and reaction rate constants.
- Specific ligand-target interactions governed by orbital-controlled mechanisms (e.g., nucleophilic or electrophilic attack).
- Investigations where the electronic energy or the distribution of electron density is a critical determinant of activity.
Prioritize topological descriptors when [20] [21]:
- Screening very large chemical databases for rapid similarity assessment or initial activity profiling.
- The biological activity is primarily influenced by molecular size, shape, or branching patterns rather than precise electronic effects.
- Computational resources or time are limited, as topological descriptors are fast and inexpensive to compute.
3. My 3D-QSAR model performance is poor. Could the molecular alignment be the issue?
Yes, the alignment of molecules is a critical step in 3D-QSAR methods like CoMFA and CoMSIA and is a common source of poor model performance. [24] To troubleshoot:
- Verify the Pharmacophore Hypothesis: Ensure alignment is based on a robust pharmacophore model that reflects the key functional groups responsible for biological activity.
- Check Conformational Selection: Confirm that all molecules are in their biologically active conformation. Using a low-energy conformation that is not the bioactive one can mislead the model.
- Use Multiple Alignment Rules: Test different alignment rules (e.g., based on a common scaffold, field fit, or receptor site) to see which yields the most predictive and interpretable model.
Troubleshooting Guides
Guide 1: Addressing Overfitting in QSAR Models During Descriptor Selection
Overfitting occurs when a model is too complex and learns noise from the training data instead of the underlying structure-activity relationship, leading to poor predictive performance on new compounds.
Protocol:
- Initial Descriptor Pool Calculation: Calculate a wide range of descriptors (e.g., using software like PaDEL-Descriptor, Dragon, or Mordred). [21] [3]
- Data Reduction: Remove descriptors with low variance or high correlation to others to reduce redundancy. [21]
- Feature Selection: Apply robust feature selection methods to identify the most relevant descriptors.
- Filter Methods: Select descriptors based on univariate statistical tests (e.g., correlation with the activity). [3]
- Wrapper Methods: Use algorithms like Genetic Algorithms to find the descriptor subset that optimizes model performance. [21]
- Embedded Methods: Utilize techniques like LASSO regression, which performs variable selection as part of the model building process. [3]
- Apply the "5:1 Rule": As a rule of thumb, have no more than one descriptor for every five compounds in the training set to maintain a good data-point-to-descriptor ratio. [21]
- Validate Rigorously:
- Internal Validation: Use k-fold cross-validation (e.g., 5-fold or 10-fold) on the training set. [3] A high cross-validated R² (Q²) is a good indicator of robustness.
- External Validation: Test the final model on a completely independent set of compounds that were not used in model building or selection. This is the gold standard for assessing predictive ability. [3]
Flowchart: A rigorous workflow to prevent overfitting during descriptor selection.
Guide 2: Selecting an Appropriate Level of Theory for Quantum Chemical Descriptors
The accuracy of quantum chemical (QC) descriptors depends on the computational method (level of theory) used. An inappropriate choice can lead to inaccurate descriptors and flawed models.
Protocol:
- Define the Requirement for Accuracy vs. Cost: Balance computational cost with the required accuracy. Density Functional Theory (DFT) is often the best compromise, offering good accuracy for reasonable cost for many systems. [22]
- Select a Functional and Basis Set: For DFT, popular general-purpose functionals include B3LYP and ωB97X-D. Pair with a basis set like 6-31G* for a good starting point. [22]
- Validate with a Test Set: For a small, representative subset of your molecules, calculate the QC descriptors using a higher level of theory (e.g., ab initio) and compare with your chosen method. If the trends are consistent, the chosen method is likely sufficient. [22]
- Consider the System: For systems with significant dispersion forces (e.g., π-π stacking), use a functional that accounts for these (e.g., ωB97X-D). For open-shell systems, use an unrestricted method. [22]
- Ensure Geometry Optimization: Always use fully optimized molecular geometries at the same level of theory used for descriptor calculation. Using non-optimized or poorly optimized structures is a common error. [22]
Table 2: Troubleshooting Quantum Chemical Descriptor Calculations
| Problem | Potential Cause | Solution |
|---|---|---|
| Unphysically high energy values | Incorrect electronic state specification (e.g., singlet vs. triplet). | Re-check the multiplicity and charge of the molecule. |
| Descriptors fail to correlate with activity | Level of theory is inadequate; descriptors are inaccurate. | Re-calculate with a higher level of theory (e.g., larger basis set, different functional). |
| Calculation fails to converge | Molecular geometry is unstable or has symmetry issues. | Tweak the initial geometry or use a computational software's built-in stability analysis. |
| Long computation times for large molecules | Using high-level ab initio methods on large, flexible molecules. | Switch to DFT or a well-parameterized semi-empirical method (e.g., PM7). [22] |
Guide 3: Implementing a Robust 3D-QSAR Workflow with CoMFA/CoMSIA
A systematic workflow is essential for building interpretable and predictive 3D-QSAR models.
Protocol:
- Data Set Preparation: Curate a set of molecules with known biological activities and defined stereochemistry. [3]
- Molecular Construction and Optimization: Build 3D structures and optimize their geometry using molecular mechanics (e.g., with MMFF94) or quantum chemical methods. [24]
- Conformational Analysis: For flexible molecules, determine the likely bioactive conformation, often the global energy minimum or a conformation aligned with a known rigid inhibitor. [24]
- Molecular Alignment: This is the most critical step. Align molecules based on a common scaffold or a pharmacophore hypothesis. This defines their relative orientation in the 3D grid. [24]
- Descriptor Generation (Field Calculation):
- Place the aligned molecules into a 3D grid.
- For CoMFA, calculate steric (Lennard-Jones) and electrostatic (Coulombic) interaction energies between a probe atom and each molecule at every grid point. [24]
- For CoMSIA, similar fields are calculated but with a Gaussian function, avoiding singularities and making results less sensitive to molecular alignment. [24]
- Partial Least Squares (PLS) Analysis: Use PLS regression to correlate the field values (descriptors) with the biological activity. [24] [3]
- Model Validation and Visualization: Validate the model via cross-validation and an external test set. Visualize the results as 3D coefficient contour maps, showing regions where steric or electrostatic changes are favorable or unfavorable for activity. [24]
Flowchart: Key steps for building a 3D-QSAR model with CoMFA/CoMSIA.
The Scientist's Toolkit: Essential Research Reagents & Software
Table 3: Essential Software Tools for Descriptor Calculation and QSAR Modeling
| Tool Name | Type/Function | Key Utility |
|---|---|---|
| Dragon | Software | Calculates thousands of molecular descriptors (2D/3D). Industry standard for comprehensive descriptor profiles. [21] |
| PaDEL-Descriptor | Software | An open-source alternative for calculating 2D and 1D molecular descriptors. [3] |
| Gaussian, GAMESS | Software | Perform quantum chemical calculations to derive accurate quantum chemical descriptors (HOMO, LUMO, etc.). [22] |
| Multiwfn | Software | A powerful wavefunction analyzer for calculating and analyzing a wide range of quantum chemical descriptors from computed wavefunctions. [22] |
| Sybyl (Tripos) | Software Suite | The commercial platform historically containing the CoMFA and CoMSIA routines for 3D-QSAR. [24] |
| RDKit | Open-Source Toolkit | A collection of cheminformatics and machine-learning software; can calculate descriptors and integrate with Python-based modeling workflows. [3] |
The Critical Link Between Descriptor Selection and Model Interpretability
Troubleshooting Guide: Common Descriptor Selection Issues
This guide addresses frequent challenges researchers face when selecting molecular descriptors for QSAR studies, impacting model interpretability and performance.
1. Problem: The "Black Box" Model
- Symptoms: Your machine learning model has good predictive power, but you cannot explain which molecular features drive the activity predictions.
- Root Cause: Over-reliance on high-dimensional, correlated descriptors that lack clear chemical meaning [25] [5].
- Solution: Implement descriptor importance adjustment methods, such as the modified Counter-Propagation Artificial Neural Network (CPANN) that dynamically weights descriptor importance during training [25]. Combine this with interpretability techniques like SHAP (SHapley Additive exPlanations) analysis to quantify each descriptor's contribution to predictions [26].
2. Problem: Model Fails to Generalize
- Symptoms: High accuracy on training data but poor performance on new compounds.
- Root Cause: Descriptor selection does not account for the model's Applicability Domain (AD), or includes redundant, highly correlated descriptors [26].
- Solution:
- Preprocessing: Remove highly correlated descriptors (e.g., Pearson’s |r| > 0.95) to reduce multicollinearity [26].
- Domain Assessment: Calculate the Mahalanobis Distance for new compounds to verify they fall within the training set's chemical space [26]. Use the χ² distribution to set a statistically valid threshold.
3. Problem: Spurious Correlations Mislead Design
- Symptoms: The model highlights "bulk" properties (e.g., molecular weight) as important, which are likely proxies for true causal features like specific pharmacophores [5].
- Root Cause: Standard machine learning models identify correlations but cannot distinguish causal relationships from confounding factors [5].
- Solution: Employ causal inference frameworks like Double/Debiased Machine Learning (DML). This method estimates the unconfounded causal effect of each descriptor by treating all others as potential confounders, providing more reliable guidance for synthesis [5].
4. Problem: Mechanistic Interpretation is Impossible
- Symptoms: Unable to relate model-selected descriptors to known toxicological mechanisms or structural alerts.
- Root Cause: Descriptors are purely statistical constructs without established links to physicochemical properties or biological mechanisms [25].
- Solution: Prioritize descriptors that can be mapped to known mechanistic features. For example, in carcinogenicity models, descriptors like
nRNNOx(number of N-nitroso groups) can be directly linked to the structural alert "alkyl and aryl–N-nitroso groups" known to form DNA adducts [25].
Frequently Asked Questions (FAQs)
Q1: What are the key criteria for selecting interpretable molecular descriptors? A descriptor should ideally be:
- Computationally Feasible: Reasonable to calculate for large compound libraries [27].
- Distinct Chemical Meaning: Linked to a understandable molecular or physicochemical property (e.g., logP, polar surface area, hydrogen bond count) [27].
- Mechanistically Relevant: Correlated with the known or proposed biological mechanism of action [25].
- Non-Redundant: Provides unique information not captured by other descriptors in the set [26].
Q2: How can I balance model complexity with interpretability? Use genetic algorithms (GA) for feature selection. The GA optimizes a fitness function (e.g., adjusted R²) that rewards model performance while penalizing complexity (number of descriptors) [26]. This inherently leads to simpler, more interpretable models without sacrificing excessive predictive power.
Q3: My model is interpretable, but my peers question its mechanistic validity. How can I address this? Adhere to the OECD's fifth principle for QSAR validation, which recommends "a mechanistic interpretation, if possible" [25]. Strengthen your interpretation by:
- Explicitly linking key descriptors from your model to established structural alerts or pharmacophores from literature [25].
- Using hypothesis testing frameworks like the Benjamini-Hochberg procedure on causal descriptor effects to control the False Discovery Rate (FDR) and provide statistical rigor [5].
Q4: What are the best practices for validating that my descriptor selection is sound?
- Internal Validation: Use rigorous cross-validation and metrics like Q² and RMSE to ensure the model is robust [26].
- External Validation: Test the model on a completely held-out test set [26].
- Domain of Applicability: Always define and report the model's applicability domain using methods like Mahalanobis Distance to clarify for which compounds the interpretations are valid [26].
Experimental Protocols for Robust Descriptor Selection
Protocol 1: Genetic Algorithm for Optimal Descriptor Selection
This methodology is used to identify a compact, optimal subset of descriptors that maximizes model performance and interpretability [26].
- Descriptor Calculation & Preprocessing: Compute a diverse set of molecular descriptors (constitutional, topological, electronic, geometrical) using software like ChemoPy or PaDEL. Standardize the resulting matrix by centering to the mean and scaling to unit variance. Remove descriptors with zero variance or excessive missing values [26].
- Correlation Filtering: Reduce multicollinearity by calculating the Pearson correlation matrix and removing one descriptor from any pair with |r| > 0.95 [26].
- Genetic Algorithm Setup:
- Representation: Use a binary chromosome where each gene represents the presence (1) or absence (0) of a specific descriptor.
- Fitness Function: Use a function like
Fitness = R²_adj - (k/n), wherekis the number of selected descriptors andnis the number of training samples. This penalizes overly complex models [26]. - Evolution: Run the GA for a set number of generations (e.g., 50) or until performance plateaus.
- Model Construction: Build a Multiple Linear Regression (MLR) model using the final subset of GA-selected descriptors. The model takes the form:
pIC50 = β₀ + β₁x₁ + β₂x₂ + … + βₙxₙ, where eachxis a selected descriptor and its coefficientβindicates the magnitude and direction of its effect on activity [26].
Protocol 2: Causal Descriptor Identification with Double Machine Learning
This advanced protocol helps distinguish causally influential descriptors from spurious correlates [5].
- Problem Formulation: Treat the assessment of each descriptor's effect as a causal inference problem, where all other descriptors are potential confounders.
- Double Machine Learning Workflow:
- For each candidate descriptor
x_i, use a machine learning model (e.g., Random Forest) to predict the biological activityyusing all other descriptors (the confounder set,z). - Simultaneously, use another ML model to predict the candidate descriptor
x_iusing the same confounder setz. - The causal effect of
x_iis estimated from the residuals of these two models, effectively "deconfounding" the relationship [5].
- For each candidate descriptor
- Hypothesis Testing: Apply the Benjamini-Hochberg procedure to the p-values of all causal effect estimates to control the False Discovery Rate (FDR). This provides a statistically sound list of descriptors with significant causal links to the activity [5].
The Scientist's Toolkit: Research Reagent Solutions
| Tool / Reagent | Function in Descriptor Selection & QSAR |
|---|---|
| ChemoPy | A Python package for calculating a comprehensive set of molecular descriptors (topological, constitutional, etc.) from chemical structures [26]. |
| Genetic Algorithm (GA) | An optimization technique used to select an optimal, minimal subset of descriptors by balancing model performance and complexity [26]. |
| SHAP (SHapley Additive exPlanations) | A game-theoretic method to interpret model predictions by quantifying the marginal contribution of each descriptor to the final prediction for any given compound [26]. |
| Double Machine Learning (DML) | A causal inference framework used to estimate the unconfounded causal effect of a descriptor on biological activity, filtering out spurious correlations [5]. |
| Mahalanobis Distance | A statistical measure used to define the Applicability Domain of a QSAR model, identifying compounds that are too dissimilar from the training set for reliable prediction [26]. |
Workflow Visualization
The diagram below illustrates a robust workflow for selecting interpretable molecular descriptors, integrating key steps from the troubleshooting guides and experimental protocols.
Comparative Analysis of QSAR Modeling Algorithms
The table below summarizes the performance and interpretability of different machine learning algorithms used in QSAR modeling, as demonstrated in a study on KRAS inhibitors [26].
| Modeling Algorithm | Key Advantage | Interpretability Strength | Example Performance (R²) [26] |
|---|---|---|---|
| Partial Least Squares (PLS) | Handles multicollinearity well via latent variables. | Good; variable importance in projection (VIP) scores indicate descriptor relevance. | 0.851 |
| Genetic Algorithm-MLR (GA-MLR) | Optimally balances model size and predictive power. | High; produces a simple, transparent linear equation with defined coefficients. | 0.677 |
| Random Forest (RF) | Robust to overfitting and noise. | Moderate; provides permutation-based importance and is compatible with SHAP analysis. | 0.796 |
| XGBoost | High predictive accuracy with complex data. | Moderate; compatible with SHAP for non-linear effect interpretation. | Not Specified |
From Theory to Practice: Strategic Descriptor Selection and Implementation in Modern QSAR
Frequently Asked Questions
Q1: My QSAR model is sensitive to outliers in the biological activity data. Which variable selection method should I use? The LAD-LASSO (Least Absolute Deviation-Least Absolute Shrinkage and Selection Operator) is specifically designed to handle this issue. Unlike standard LS-LASSO, which uses a least squares criterion sensitive to outliers, LAD-LASSO employs a least absolute deviation criterion that is robust against heavy-tailed errors and severe outliers [28]. This method provides low bias in estimating large coefficients and maintains good prediction performance even when outlier observations are present in your dataset [28].
Q2: How does the choice of mutual information estimator impact the performance of the mRMR feature selection method? The performance of the Maximum Relevance Minimum Redundancy (mRMR) method is highly dependent on the mutual information estimator chosen. Different estimators, such as the Parzen window, equidistant partitioning (cells method), or bias-corrected versions, can yield varying results [29]. The estimator must be carefully selected based on your dataset characteristics, as an inappropriate choice can lead to unreliable feature selection. A bias-corrected estimator often improves mRMR performance by providing more stable mutual information assessments [29].
Q3: When should I prefer mutual information-based methods over Genetic Algorithms for variable selection in QSAR studies? Mutual information methods are generally preferred when you need computational efficiency and want to capture both linear and nonlinear dependencies between descriptors and biological activity [29] [30]. Genetic Algorithms are more appropriate when you're exploring a complex feature space and want to avoid local minima, though they can be computationally intensive [31]. For high-dimensional descriptor spaces, mutual information methods like mRMR or DMIM often provide better computational performance [29] [30].
Q4: What are the key differences between filter methods (like mutual information) and wrapper methods (like Genetic Algorithms)? Filter methods (e.g., mutual information) evaluate features based on intrinsic data properties, independent of a specific classifier, making them computationally efficient and model-agnostic [29]. Wrapper methods (e.g., Genetic Algorithms) use the performance of a specific predictive model to evaluate feature subsets, potentially yielding better performance but at higher computational cost and with potential overfitting risks [31]. Embedded methods like LASSO incorporate feature selection directly into the model training process, providing a balance between both approaches [28].
Q5: Why does my LASSO-selected model show high bias in estimating large coefficients? This is a known limitation of standard LS-LASSO (Least Squares-LASSO), which can produce high bias when estimating large coefficients [28]. Consider using robust variants like LAD-LASSO, which demonstrates lower bias for large coefficient estimation while maintaining the sparsity and variable selection capabilities of traditional LASSO [28]. The bias arises from the simultaneous variable selection and parameter estimation in LS-LASSO, which LAD-LASSO mitigates through its robust objective function [28].
Troubleshooting Guides
Issue 1: Poor Generalization Performance Despite High Training Accuracy
Problem: Your QSAR model performs well on training data but poorly on external test sets after variable selection.
Solution:
- Apply stricter validation protocols: Use external test sets that were completely excluded from the variable selection process [32].
- Evaluate applicability domain: Ensure test compounds fall within the chemical space of your training set [28] [27].
- Check for overfitting in selection: Use cross-validation during variable selection, not just during model building [32].
- Consider simpler models: If using Genetic Algorithms, they may overfit with too many generations; reduce population size or generations [31].
Performance metrics to check:
- Concordance Correlation Coefficient (CCC) should be >0.8 for external validation [32]
- rm² metric should meet established thresholds [32]
- R² and MSE for both training and test sets should be comparable [28]
Issue 2: Inconsistent Variable Selection Across Different Algorithms
Problem: Different variable selection methods (GA, LASSO, Mutual Information) yield different descriptor subsets for the same dataset.
Solution:
- Analyze descriptor redundancy: High correlation between descriptors can cause this issue. Pre-filter strongly correlated descriptors (e.g., |r| > 0.95) [28].
- Check statistical significance: Use multiple methods and select descriptors consistently identified across methods [31].
- Evaluate stability: Use bootstrap resampling to assess selection frequency of each descriptor [28].
- Prioritize interpretability: Select the descriptor set that aligns best with known structure-activity relationships in your chemical domain [27].
Issue 3: Computational Limitations with High-Dimensional Descriptor Spaces
Problem: Variable selection becomes computationally prohibitive with thousands of molecular descriptors.
Solution:
- Implement pre-filtering: Use simple correlation measures or variance thresholds to reduce descriptor space before applying advanced methods [28].
- Choose efficient algorithms: For high-dimensional spaces, mRMR with efficient mutual information estimators often outperforms Genetic Algorithms computationally [29].
- Use embedded methods: LASSO variants perform selection during modeling, reducing computational overhead compared to wrapper methods [28].
- Leverage optimized software: Use specialized packages like DRAGON with built-in selection tools rather than custom implementations [28].
Performance Comparison of Variable Selection Methods
Table 1: Key Characteristics of Variable Selection Methods in QSAR
| Method | Key Strengths | Limitations | Optimal Use Cases |
|---|---|---|---|
| Genetic Algorithms | Effective for complex feature spaces; Avoid local minima [31] | Computationally intensive; Risk of overfitting [31] | Moderate-dimensional data (<500 descriptors); Complex nonlinear relationships [31] |
| LASSO/LAD-LASSO | Simultaneous selection & estimation; Robust to outliers (LAD-LASSO) [28] | High bias for large coefficients (standard LASSO) [28] | High-dimensional data; When interpretability is important [28] |
| Mutual Information (mRMR) | Captures nonlinear dependencies; Computationally efficient [29] | Performance depends on estimator choice [29] | Large datasets; When both linear & nonlinear relationships exist [29] |
| Decomposed Mutual Information (DMIM) | Overcomes complementarity penalization [30] | Less established in QSAR literature [30] | Classification tasks; When complementary features are important [30] |
Table 2: Typical Performance Metrics for Validated QSAR Models
| Validation Type | Metric | Acceptable Threshold | Notes |
|---|---|---|---|
| Internal | Q² (LOO-CV) | >0.5 | May be optimistic [32] |
| External | R²test | >0.6 | Should be close to R²training [28] [32] |
| External | CCC | >0.8 | More reliable than R² alone [32] |
| External | rm² | >0.5 | Specific for QSAR validation [32] |
| Overall | MSEtest | As low as possible | Should be comparable to MSEtraining [28] |
Experimental Protocols
Protocol 1: Implementing LAD-LASSO for Robust Variable Selection
Purpose: Select molecular descriptors while maintaining robustness to outliers in biological activity data.
Materials:
- Dataset with molecular structures and biological activities
- DRAGON software or equivalent for descriptor calculation [28]
- MATLAB, R, or Python with optimization packages
- Preprocessing tools for descriptor filtering
Procedure:
- Calculate descriptors: Compute molecular descriptors using DRAGON (≈3224 descriptors) [28]
- Preprocess descriptors:
- Remove constant and near-constant variance descriptors
- Eliminate highly correlated descriptors (|r| > 0.95) [28]
- Standardize remaining descriptors to zero mean and unit variance
- Implement LAD-LASSO optimization:
- Use the objective function: minβ(∑|yi - xi'β| + λ∑|βj|)
- Apply cross-validation to select optimal λ parameter
- Use alternating direction method of multipliers (ADMM) for efficient optimization [33]
- Select final descriptors: Choose descriptors with non-zero coefficients in the optimal model
- Validate selection: Build QSAR model with selected descriptors and evaluate using external test set
Troubleshooting Notes:
- If convergence is slow, adjust optimization tolerance parameters
- If too many/few descriptors are selected, adjust λ range in cross-validation
- Verify robustness by artificially introducing outliers and comparing with standard LASSO
Protocol 2: mRMR with Bias-Corrected Mutual Information Estimation
Purpose: Select features with maximum relevance to activity and minimum redundancy among themselves using improved mutual information estimation.
Materials:
- Dataset with continuous molecular descriptors and biological activity
- Programming environment with mutual information estimation capabilities
- mRMR implementation with customizable estimator options
Procedure:
- Prepare data: Preprocess descriptors and ensure continuous format
- Select mutual information estimator: Choose from:
- Parzen window estimation
- Equidistant partitioning (cells method)
- Bias-corrected estimator (recommended) [29]
- Implement mRMR algorithm:
- Initialize selected feature set S = ∅
- For each iteration:
- Calculate relevance: rel(f) = I(f; C) for all features f ∉ S
- Calculate redundancy: red(f) = (1/|S|) ∑ I(f; s) for all s ∈ S
- Select feature maximizing: rel(f) - red(f)
- Add to S [29]
- Add regularization (optional): Include small regularization term in denominator for numerical stability [29]
- Determine stopping criterion: Use predefined number of features or performance plateau
Validation:
- Compare with other estimators to assess stability
- Evaluate final feature set using classification/regression performance
- Use Y-randomization to confirm significance [28]
The Scientist's Toolkit
Table 3: Essential Computational Tools for Variable Selection in QSAR
| Tool/Software | Primary Function | Application in Variable Selection |
|---|---|---|
| DRAGON | Molecular descriptor calculation | Calculates 3224+ molecular descriptors for QSAR [28] |
| Mordred | Molecular descriptor calculation | Python package for calculating 1600+ molecular descriptors [34] |
| MATLAB | Numerical computing | Implementation of LAD-LASSO and correlation filtering [28] |
| PaDEL-Descriptor | Molecular descriptor calculation | Generates molecular descriptors for cheminformatics [3] |
| Cerius2 | Molecular modeling | Includes Genetic Algorithm (GFA) for variable selection [31] |
| MOE | Molecular modeling | QuaSAR-Evolution module for GA-based selection [31] |
Workflow Visualization
In Quantitative Structure-Activity Relationship (QSAR) research, the initial set of calculated molecular descriptors is often vast and highly dimensional. Datasets can contain hundreds to thousands of descriptors, many of which are redundant, noisy, or irrelevant for predicting biological activity. This high dimensionality poses significant challenges, including overfitting, increased computational costs, and difficulty in model interpretation—a phenomenon known as the "curse of dimensionality" [35] [36]. Dimensionality reduction techniques are therefore not merely optional pre-processing steps but are fundamental to developing robust, interpretable, and predictive QSAR models. This technical support guide focuses on two powerful, complementary methods: Principal Component Analysis (PCA), a feature extraction technique, and Recursive Feature Elimination (RFE), a feature selection method. We provide detailed troubleshooting and FAQs to help researchers effectively implement these techniques within their QSAR workflows.
The table below summarizes the key characteristics of PCA and RFE to help you select the appropriate strategy.
| Feature | Principal Component Analysis (PCA) | Recursive Feature Elimination (RFE) |
|---|---|---|
| Category | Feature Extraction [35] | Feature Selection [35] [37] |
| Core Principle | Projects data to a new, lower-dimensional space of orthogonal Principal Components (PCs) that maximize variance [35] [36]. | Iteratively removes the least important features based on a model's feature importance scores [37]. |
| Output | Principal Components (PCs)—linear combinations of all original features [35]. | A subset of the original, interpretable molecular descriptors [37]. |
| Interpretability | Low; PCs are mathematical constructs and often lack direct chemical meaning [35]. | High; retains the original descriptors, allowing for direct structure-activity interpretation [37]. |
| Primary Use Case | Dealing with multicollinearity; reducing noise; visualizing high-dimensional data [38] [39]. | Identifying the most impactful molecular descriptors to guide lead optimization [37]. |
Experimental Protocols and Workflows
Protocol 1: Implementing PCA for Descriptor Reduction
PCA is an unsupervised technique from linear algebra used to project a dataset into a lower-dimensional space while preserving its essential variance [35] [36].
Detailed Methodology:
- Data Standardization: Before applying PCA, standardize your descriptor matrix so that each descriptor has a mean of 0 and a standard deviation of 1. This ensures all descriptors contribute equally to the variance, regardless of their original units [35].
- Covariance Matrix Computation: Calculate the covariance matrix of the standardized data to understand how the descriptors vary with one another.
- Eigendecomposition: Perform eigendecomposition on the covariance matrix to obtain eigenvalues and eigenvectors. The eigenvectors represent the principal components (PCs), and the eigenvalues indicate the amount of variance captured by each PC [35].
- Projection: Project the original data onto the selected principal components to create a new, reduced dataset [36].
Key Considerations:
- Number of Components: The choice of how many PCs to retain is critical. A common approach is to select the number that captures a pre-defined threshold of the total variance (e.g., 90-95%). This can be determined by analyzing the scree plot, which plots the eigenvalues in descending order [36].
- Variance Threshold: Retain components that contribute significantly to the total variance and discard those with negligible contribution.
Protocol 2: Implementing RFE for Feature Selection
RFE is a supervised wrapper feature selection method that recursively prunes the least important features from a model to find the optimal subset that maximizes predictive performance [37].
Detailed Methodology:
- Model Training: Train a supervised learning algorithm (e.g., Random Forest or Support Vector Machine) capable of outputting feature importance scores on the entire set of descriptors.
- Feature Ranking: Rank all molecular descriptors based on the model's feature importance scores (e.g.,
feature_importances_in scikit-learn) [37]. - Feature Pruning: Remove the least important feature(s) from the current set.
- Recursion: Repeat steps 1-3 on the progressively smaller subset of descriptors until the desired number of features is reached.
- Performance Evaluation: At each iteration, evaluate the model's performance (e.g., via cross-validated accuracy or R²) to identify the subset of features that yields the best performance [37].
Key Considerations:
- Base Model: The choice of the underlying model (e.g., Random Forest) is crucial as it determines how feature importance is calculated.
- Optimal Feature Count: The final number of features to select is a hyperparameter. It is determined by identifying the point where model performance is maximized before it begins to degrade due to the removal of important features [37].
The Scientist's Toolkit: Essential Research Reagents & Software
The table below lists key computational tools and resources essential for implementing PCA and RFE in QSAR studies.
| Tool/Resource | Function | Application in QSAR |
|---|---|---|
| PaDEL-Descriptor [40] [3] | Calculates molecular descriptors and fingerprints. | Generates the initial high-dimensional feature set from chemical structures. |
| scikit-learn (Python) [36] | Machine learning library containing PCA, RFE, and various estimators. | Provides the primary API for implementing the dimensionality reduction protocols. |
| R Statistical Environment [40] | Platform for statistical computing and graphics. | Used for model building, validation, and generating dynamic analysis reports. |
| KNIME / RapidMiner [40] [41] | Graphical workflow platforms for data analytics. | Enables the construction of reproducible, visual pipelines for QSAR modeling. |
| Dragon [41] [3] | Commercial software for calculating a wide range of molecular descriptors. | An alternative to PaDEL for comprehensive descriptor calculation. |
Troubleshooting Guides & FAQs
Frequently Asked Questions
Q1: My QSAR model's performance dropped after applying PCA. What could be the cause? This often occurs when biologically relevant variance is not the primary variance captured by the first few principal components. PCA is unsupervised and selects components that maximize total variance in the descriptor space, which may not always align with variance predictive of the biological activity. Consider using supervised dimensionality reduction methods or applying feature selection (like RFE) instead.
Q2: How do I determine the optimal number of features to select with RFE? The optimal number is not predetermined. You must perform RFE iteratively, evaluating model performance (e.g., using cross-validated accuracy or R²) at each step. Plot the performance metric against the number of features. The optimal number is typically at or near the point of peak performance before it starts to decline [37].
Q3: Can PCA and RFE be used together in a QSAR workflow? Yes, this is a powerful and common strategy. You can use PCA initially to reduce noise and handle multicollinearity among a large number of descriptors. The resulting PCs can then be used in an RFE process to further refine the most predictive components, although this sacrifices some interpretability.
Q4: Why are my selected molecular descriptors from RFE chemically unintelligible or difficult to interpret? Some powerful molecular descriptors (e.g., certain topological or quantum chemical indices) are inherently complex. Focus on identifying the physicochemical properties these descriptors represent (e.g., lipophilicity, polarity, molecular size). This abstraction can provide the chemical insight needed to guide molecular design [39].
Common Issues and Solutions
| Problem | Potential Cause | Solution |
|---|---|---|
| PCA results are dominated by a few descriptors. | Descriptors were not standardized before applying PCA, so those with larger scales dominate the variance. | Always standardize data (mean=0, std=1) before performing PCA [35]. |
| RFE is computationally slow for a large descriptor set. | The base model is being retrained a very large number of times. | Increase the number of features removed per step. Use a faster base model or perform an initial filter-based feature selection to reduce the starting set. |
| Model performance is unstable after RFE. | The selected feature subset is too small or sensitive to small changes in the training data. | Use a more robust model like Random Forest as the RFE estimator. Use repeated cross-validation to get a more stable estimate of performance for each subset. |
| Poor external validation performance after dimensionality reduction. | The applicability domain of the model has been violated, or the reduction was overfitted to the training set. | Ensure the PCA transformation or RFE feature set is derived only from the training data and then applied to the test set. Define and check the applicability domain of your final model [3]. |
Troubleshooting Guide: Descriptor Selection in QSAR Modeling
FAQ: My QSAR model performs well on the training data but poorly on new compounds. What is the issue? This is a classic sign of overfitting, where the model has memorized the training data noise instead of learning the generalizable structure-activity relationship. To resolve this:
- Apply Rigorous Feature Selection: Reduce the number of descriptors. In a study on NF-κB inhibitors, researchers started with 17,967 descriptors. They applied a Pearson correlation-based filter (cutoff of 0.6) to remove highly correlated features, followed by univariate analysis and SVC-L1 regularization to select the most statistically significant descriptors, ultimately reducing the feature set to a more robust size [42].
- Define the Applicability Domain (AD): Use methods like the leverage method to define the chemical space your model is valid for. Predictions for compounds outside this domain are unreliable [43].
- Simplify the Model: If your dataset is limited, a simpler linear model (e.g., MLR or PLS) may generalize better than a complex non-linear model (e.g., ANN) [3].
FAQ: How many molecular descriptors should I use for my model? There is no fixed number, but the ratio of compounds to descriptors should be sufficient to avoid chance correlations. Best practices involve:
- Use Feature Selection Algorithms: Do not rely on a arbitrary number. Employ filter methods (like correlation analysis), wrapper methods (like genetic algorithms), or embedded methods (like LASSO) to identify the optimal descriptor subset [3].
- Prioritize Interpretability: A model with fewer, chemically meaningful descriptors is often more valuable than a "black box" with thousands. The case study on 121 NF-κB inhibitors successfully developed a simplified MLR model with a reduced number of terms that maintained accuracy [43].
FAQ: What types of descriptors are most informative for modeling NF-κB inhibition? Research indicates that a combination of descriptor types is effective.
- 2D and Fingerprint Descriptors: For classifying TNF-α induced NF-κB inhibitors, models built using 2D descriptors and molecular fingerprints achieved higher predictive accuracy (AUC up to 0.66) than those using 3D descriptors alone (AUC 0.56) [42].
- Steric and Hydrophobic Properties: Although for a different target (Histamine H3R), QSAR models consistently showed that steric and hydrophobic properties of ligands are critical for good biological affinity [44]. This highlights the importance of these fundamental physicochemical properties in drug-target interactions.
FAQ: How can I validate that my descriptor selection process is sound? Robust validation is key to a reliable QSAR model.
- Internal Validation: Use cross-validation techniques (e.g., 5-fold or 10-fold) on your training set to assess model stability [3].
- External Validation: The gold standard is to test the model on a completely independent set of compounds that were not used in model building or feature selection. In the NF-κB case study, the dataset was split into a training set (~80%) and an independent test set (~20%) for final evaluation [43] [42].
- Use of Multiple Algorithms: Compare models built using different algorithms (e.g., MLR vs. ANN) on the same selected descriptors. In one study, an ANN model demonstrated superior reliability and prediction for NF-κB inhibitors compared to an MLR model [43].
Experimental Protocol: QSAR Modeling for NF-κB Inhibitors
The following workflow is compiled from successful case studies on NF-κB inhibitor prediction [43] [42].
1. Dataset Curation
- Source: Collect a dataset of chemical compounds with experimentally determined NF-κB inhibitory activities (e.g., IC50 values). Public repositories like PubChem BioAssay (e.g., AID 1852) are common sources [42].
- Preparation: Standardize chemical structures (e.g., remove salts, normalize tautomers). Divide the dataset randomly into a training set (~80%) for model development and a test set (~20%) for external validation. Ensure both sets cover a diverse chemical space.
2. Molecular Descriptor Calculation and Preprocessing
- Calculation: Use software like PaDEL-Descriptor or Dragon to calculate a comprehensive set of 1D, 2D, and 3D molecular descriptors from the compounds' SMILES representations. This can generate thousands of initial descriptors [42].
- Preprocessing: Normalize descriptor values (e.g., using Standard Scaler for z-score normalization). Remove descriptors with a high percentage (e.g., >80%) of null values or with low variance across the dataset [42].
3. Descriptor Selection
- Correlation Filtering: Apply a Pearson correlation filter (e.g., threshold of 0.6) to remove highly correlated descriptors and reduce multicollinearity [42].
- Advanced Selection: Use statistical and machine learning-based methods to identify the most significant descriptors. Proven techniques include:
4. Model Building and Validation
- Algorithm Selection: Build models using both linear (e.g., Multiple Linear Regression - MLR) and non-linear (e.g., Artificial Neural Networks - ANN, Support Vector Machines - SVM) algorithms.
- Internal Validation: Perform k-fold cross-validation (e.g., 5-fold) on the training set to tune model parameters and prevent overfitting.
- External Validation: Use the held-out test set for the final evaluation of the model's predictive performance. Key metrics include R², Mean Absolute Error (MAE), and for classification, Area Under the Curve (AUC).
The following diagram summarizes the experimental workflow for building a validated QSAR model.
Quantitative Data from NF-κB QSAR Case Studies
The table below summarizes key data on model performance and descriptor selection from published studies.
| Study Focus | Initial Descriptors | Selected Descriptors | Best Model Performance | Key Descriptor Selection Methods |
|---|---|---|---|---|
| NF-κB Inhibitor Prediction (121 compounds) [43] | Not Specified | Reduced set of significant terms | ANN model showed superior reliability and prediction vs. MLR | Analysis of Variance (ANOVA), Leverage method for Applicability Domain |
| NF-κB Inhibitor Classification (2,481 compounds) [42] | 17,967 (1D, 2D, 3D, Fingerprints) | 2,365 post-correlation filter | Support Vector Classifier (AUC: 0.75) | Variance Threshold, Pearson Correlation (cutoff 0.6), Univariate analysis, SVC-L1 regularization |
Signaling Pathway: NF-κB Activation
Understanding the biological context is crucial for rational descriptor selection. NF-κB activation primarily occurs via the canonical pathway, which is a key target for therapeutic inhibition.
The Scientist's Toolkit: Research Reagent Solutions
| Tool / Reagent | Function in QSAR Modeling |
|---|---|
| PaDEL-Descriptor | Open-source software for calculating a wide range of 1D, 2D, and 3D molecular descriptors and fingerprints from chemical structures [42]. |
| PubChem BioAssay | A public repository providing bioactivity data for millions of compounds, serving as a primary source for curating training and test datasets [42]. |
| Standard Scaler (e.g., Scikit-learn) | A preprocessing tool used to normalize molecular descriptor values by centering (zero mean) and scaling (unit variance), ensuring descriptors contribute equally to the model [42]. |
| Support Vector Classifier (SVC) with L1 Regularization | A machine learning algorithm used not only for modeling but also for feature selection, as it can drive the coefficients of non-informative descriptors to zero [42]. |
| Artificial Neural Networks (ANNs) | A powerful non-linear modeling algorithm capable of capturing complex relationships between molecular descriptors and biological activity, often showing superior performance [43]. |
Selecting appropriate software and molecular descriptors is a foundational step in developing robust Quantitative Structure-Activity Relationship (QSAR) models. This technical support center provides a comparative overview and troubleshooting guide for four widely used computational tools—VEGA, EPI Suite, DRAGON, and ADMETLab—to assist researchers in making informed decisions for their drug discovery and environmental chemistry workflows.
The following diagram illustrates a general workflow for integrating these tools into a QSAR research project, from initial compound input to final model interpretation.
Tool Comparison & Selection Guide
The table below summarizes the core characteristics, strengths, and limitations of each software tool to guide your selection process.
| Software | Primary Function | Descriptor Types | Key Applications | Regulatory Acceptance | Access |
|---|---|---|---|---|---|
| VEGA | QSAR models & read-across | Various, model-dependent | Toxicity, environmental fate, PBT assessment | High (REACH, CLP compliant) [45] [46] | Free, platform-dependent [47] |
| EPI Suite | Physical/chemical property estimation | Fragment-based, group contribution | Environmental fate, biodegradation, bioaccumulation [48] | High (US EPA, REACH) [46] | Free, Windows-based [48] |
| DRAGON | Molecular descriptor calculation | 1D-3D structural descriptors [41] | Descriptor generation for custom QSAR, drug design | Research use | Commercial |
| ADMETLab | ADMET property prediction | 2D, fingerprints, ECFP [49] | Drug-likeness, absorption, distribution, metabolism, excretion, toxicity [49] | Research use | Free web server [49] |
Frequently Asked Questions & Troubleshooting
General Tool Selection
Q1: Which tool is most appropriate for regulatory submissions under REACH?
Both VEGA and EPI Suite are widely accepted for regulatory submissions. VEGA is particularly valuable because it provides detailed applicability domain assessment and combines QSAR with read-across approaches, which is important for REACH compliance [45] [47]. EPI Suite is recognized by US EPA and frequently used for predicting physicochemical and environmental fate properties [48] [46].
Q2: How do I handle discrepancies between predictions from different tools?
First, check the Applicability Domain (AD) of each model. VEGA provides a unique Applicability Domain Index that evaluates similarity to training compounds, descriptor ranges, and consistency with experimental data of similar compounds [47]. Prioritize predictions that fall within well-defined applicability domains. Use a weight-of-evidence approach by considering results from multiple tools and available experimental data [46] [47].
Technical Issues & Methodology
Q3: What should I do if EPI Suite fails to run or produces inconsistent structures?
The downloadable version of EPI Suite (v4.11) has known technical difficulties. The US EPA recommends using the web-based beta version (EPI Suite Beta 1.0) as an alternative [48]. If structures are interpreted inconsistently, ensure you're using standardized SMILES strings, as the EPA is implementing automatic standardization to address this issue [48].
Q4: How can I improve the reliability of VEGA predictions?
Carefully evaluate all elements provided in the VEGA report:
- Check the similarity score of related compounds (values < 0.75 indicate important structural differences) [47]
- Review the presence of structural alerts and any unusual fragments [47]
- Assess the concordance between predicted values and experimental data of similar compounds [47]
- When multiple models are available for the same endpoint, evaluate each independently and integrate results [47]
Q5: Which tool provides the most comprehensive descriptor calculation for custom QSAR models?
DRAGON specializes in calculating a wide range of molecular descriptors (1D, 2D, and 3D) for building custom QSAR models [41]. For researchers focusing on ADMET properties, ADMETLab uses robust QSAR models based on 2D descriptors and various fingerprints (MACCS, ECFP) with published performance metrics [49].
Experimental Protocol: Tool Integration Workflow
For comprehensive chemical assessment, follow this integrated protocol:
- Input Standardization: Generate standardized SMILES or MOL files for all compounds using tools like RDKit [50].
- Descriptor Calculation (DRAGON): Calculate comprehensive molecular descriptors for custom model development [41].
- Environmental Fate Screening (EPI Suite): Estimate biodegradation (BIOWIN), bioaccumulation (BCFBAF), and soil adsorption (KOCWIN) potential [45] [48].
- Toxicity & Regulatory Assessment (VEGA): Evaluate mutagenicity, ecotoxicity, and other endpoints using multiple models. Carefully review Applicability Domain Index for each prediction [45] [47].
- ADMET Profiling (ADMETLab): Predict drug-likeness, permeability, metabolic stability, and toxicity endpoints using validated models [49].
- Results Integration: Compare predictions across tools, prioritizing those with strong applicability domain coverage and consensus.
Research Reagent Solutions
The table below outlines the essential computational "reagents" for QSAR studies and their specific functions in the research workflow.
| Tool/Resource | Function in Research | Key Outputs | Considerations |
|---|---|---|---|
| VEGA Platform | Regulatory-focused toxicity prediction | Mutagenicity, bioaccumulation, persistence predictions with reliability assessment [45] | Always check Applicability Domain Index [47] |
| EPI Suite | Environmental fate profiling | Log Kow, biodegradation probability, hydrolysis rates [48] | Use web-based beta if technical issues arise [48] |
| DRAGON | Molecular descriptor calculation | 1D, 2D & 3D molecular descriptors for custom models [41] | Commercial license required |
| ADMETLab | Comprehensive ADMET screening | 30+ ADMET endpoints including solubility, permeability, metabolism [49] | Free web server with batch computation [49] |
| Standardized SMILES | Chemical structure representation | Consistent structure interpretation across tools | Essential for reproducible results [48] [50] |
| Applicability Domain Assessment | Prediction reliability evaluation | Quantitative measures of prediction uncertainty | Critical for regulatory acceptance [45] [47] |
Key Technical Considerations
When selecting molecular descriptors for QSAR research, consider that each tool employs different descriptor strategies: EPI Suite primarily uses fragment-based methods [48], ADMETLab utilizes 2D descriptors and fingerprints [49], while DRAGON provides the most comprehensive 1D-3D descriptor calculation [41]. For regulatory applications, complement descriptor-based predictions with VEGA's read-across capabilities and always verify predictions fall within the model's applicability domain to ensure reliability [45] [47].
Balancing Interpretability and Predictive Power in Descriptor Selection
Frequently Asked Questions
What is the most common pitfall when selecting molecular descriptors for a QSAR model? The most common pitfall is high information redundancy among descriptors, where strongly correlated descriptors can constitute over 90% of the initial descriptor pool. This redundancy can lead to model overfitting and reduced interpretability without improving predictive power. A Representative Feature Selection (RFS) approach that calculates Euclidean distances and Pearson correlation coefficients can effectively reduce this redundancy and enhance model performance [51].
How can I improve my QSAR model's interpretability without sacrificing predictive accuracy? Utilize methods that provide inherent interpretability, such as Genetic Algorithm-optimized Multiple Linear Regression (GA-MLR), which selects an optimal subset of descriptors while maintaining a transparent linear model structure. These models can achieve robust predictive performance (R² = 0.677 in KRAS inhibitor studies) while remaining chemically interpretable. Additionally, SHapley Additive exPlanations (SHAP) can be applied to any model for prediction-wise feature importance analysis [26].
My QSAR model performs well on training data but poorly on new compounds. What might be wrong? This typically indicates an applicability domain issue. Your model may be making predictions for compounds structurally different from its training set. Implement applicability domain assessment using methods like Mahalanobis Distance with a threshold based on the 95th percentile of the χ² distribution. This helps flag compounds outside the domain where your model can reliably predict [26].
What are the practical trade-offs between traditional molecular descriptors and deep learning representations? Traditional descriptors (e.g., topological, constitutional, electronic) offer better interpretability and direct chemical insights but may require careful feature selection to avoid redundancy. Deep learning representations can automatically extract relevant features and sometimes achieve higher predictive performance (R² up to 0.92 in advanced Bio-QSARs) but operate as "black boxes" requiring additional interpretation techniques like attention mechanisms or Layer-wise Relevance Propagation [52] [53].
How can I handle "activity cliffs" where similar structures have very different activities? Similarity-based methods like Metric Learning Kernel Regression (MLKR) or Topological Regression can address this by learning a supervised similarity metric that incorporates activity information. These techniques create smoother activity landscapes where chemically-similar-but-functionally-different molecules are properly separated in the representation space [53].
Troubleshooting Guides
Problem: Poor Model Performance Despite Extensive Descriptor Pool
Symptoms:
- Low R² values on test set (< 0.6)
- High root mean square error (RMSE)
- Large discrepancy between training and test performance
Solution Steps:
- Apply rigorous feature selection: Use Genetic Algorithms or Representative Feature Selection to identify an optimal descriptor subset [51] [26].
- Reduce multicollinearity: Remove highly correlated descriptors (Pearson's |r| > 0.95) [26].
- Expand chemical diversity: Ensure training set covers structural space of interest [27].
- Try ensemble methods: Implement Random Forest or XGBoost which can handle descriptor redundancy better [26].
Verification: After implementing RFS, model performance should show improved test set R² (> 0.7) and reduced overfitting.
Problem: Model Predictions Lack Chemical Interpretability
Symptoms:
- Inability to explain which structural features drive activity
- No clear guidance for molecular optimization
- Resistance from medicinal chemists to use predictions
Solution Steps:
- Use inherently interpretable algorithms: GA-MLR or PLS regression provide transparent models [26].
- Implement interpretation frameworks: Apply SHAP or permutation-based importance [25] [26].
- Select mechanistically-relevant descriptors: Prioritize descriptors linked to known mechanisms (e.g., logP for permeability) [27].
- Incorporate domain knowledge: Use prior knowledge to guide descriptor selection rather than purely data-driven approaches [54].
Verification: You should be able to identify specific molecular features contributing to activity and propose structural modifications with predicted effects.
Problem: Inconsistent Performance Across Different Compound Classes
Symptoms:
- Excellent predictions for some chemotypes but poor for others
- Model fails to generalize to new structural scaffolds
- High prediction variance across chemical space
Solution Steps:
- Define applicability domain: Calculate Mahalanobis distance for training set and set threshold for reliable predictions [26].
- Use similarity-based methods: Implement Topological Regression or read-across approaches [53].
- Ensure representative training data: Apply Statistical Molecular Design to cover chemical space systematically [55].
- Consider local models: Develop separate QSAR models for different chemical classes [27].
Verification: Model should provide uncertainty estimates for predictions and reliably identify when compounds are outside its expertise domain.
Problem: Balancing Simple vs. Complex Descriptors for Drug Discovery
Symptoms:
- Simple descriptors produce interpretable but inaccurate models
- Complex quantum-chemical descriptors are accurate but computationally expensive
- Difficulty communicating results to multidisciplinary teams
Solution Steps:
- Adopt a dual-solution approach: Maintain both simple and complex models for different applications [56].
- Use descriptor importance adjustment: Implement algorithms like modified CPANN that dynamically adjust descriptor importance during training [25].
- Benchmark multiple approaches: Compare traditional descriptors vs. deep learning representations for your specific endpoint [53].
- Implement hierarchical modeling: Start with simple models for rapid screening, use complex models for final candidates [27].
Verification: You should have multiple modeling approaches available with clear understanding of when to use each based on project stage and requirements.
Experimental Protocols
Protocol 1: Representative Feature Selection for Descriptor Screening
Purpose: To identify a non-redundant, representative set of molecular descriptors from a large initial pool while maintaining predictive power.
Materials and Reagents:
- Molecular structures in SMILES format
- Dragon software or equivalent descriptor calculation package
- Programming environment (Python/R) with scikit-learn
Procedure:
- Calculate 5270 molecular descriptors using Dragon software [51].
- Preprocess descriptors: remove constants, handle missing values, normalize.
- Calculate Pearson correlation coefficients between all descriptor pairs.
- Cluster descriptors using Euclidean distance and correlation threshold (|r| > 0.8).
- Within each cluster, select the descriptor with highest variance as representative.
- Validate selected descriptor set by building QSAR model and comparing performance to full descriptor set.
Expected Outcomes: Reduced descriptor set (typically 5-15% of original) with maintained or improved predictive performance (R² > 0.8 for classification tasks) [51].
Protocol 2: GA-MLR Model Development with Applicability Domain
Purpose: To develop a genetically optimized multiple linear regression QSAR model with defined applicability domain for reliable predictions.
Materials and Reagents:
- Chemical dataset with measured activity values (pIC50)
- ChemoPy or RDKit for descriptor calculation
- Genetic algorithm implementation (Python DEAP library)
- Validation framework with cross-validation
Procedure:
- Compile dataset of 62-100 compounds with standardized activity measurements [26].
- Calculate molecular descriptors (topological, constitutional, electronic).
- Preprocess descriptors: remove correlated features (|r| > 0.95), standardize.
- Implement genetic algorithm with binary chromosome representation:
- Population size: 100
- Generations: 50
- Fitness function: adjusted R² - k/n (k=descriptors, n=samples)
- Crossover probability: 0.8
- Mutation probability: 0.1
- Train multiple linear regression model with GA-selected descriptors.
- Define applicability domain using Mahalanobis distance with χ² threshold (95%).
- Validate model using 5-fold cross-validation and external test set.
Expected Outcomes: Interpretable linear model with 5-10 descriptors, good predictive performance (R² > 0.65), and clear applicability domain definition [26].
Research Reagent Solutions
| Reagent Type | Specific Examples | Function in QSAR Studies |
|---|---|---|
| Descriptor Software | Dragon, PaDEL, Mordred, ChemoPy | Calculates molecular descriptors from chemical structures [51] [26] [53] |
| Feature Selection | Genetic Algorithms, RFS, Stepwise MLR | Identifies optimal descriptor subsets, reduces redundancy [51] [26] |
| Modeling Algorithms | PLS, Random Forest, XGBoost, CPANN | Builds predictive relationships between descriptors and activity [25] [26] |
| Interpretation Tools | SHAP, Permutation Importance, Layer-wise Relevance Propagation | Explains model predictions and descriptor contributions [25] [26] [53] |
| Validation Methods | Cross-validation, Applicability Domain, Y-randomization | Ensures model robustness and reliability for new compounds [26] [55] |
Performance Comparison of Descriptor Selection Methods
| Method | Typical Descriptor Reduction | Interpretability | Predictive Performance | Best Use Cases |
|---|---|---|---|---|
| Representative Feature Selection | 85-95% reduction | High | R² ~0.8-0.9 | Large descriptor pools, classification tasks [51] |
| Genetic Algorithm-MLR | 90-98% reduction | High | R² ~0.65-0.85 | Lead optimization, mechanistic studies [26] |
| Deep Learning | Automatic feature extraction | Low | R² ~0.8-0.92 | Complex endpoints, large datasets [52] [53] |
| Topological Regression | Varies based on similarity metric | Medium-High | Competitive with deep learning | Activity cliffs, lead optimization [53] |
| PLS Regression | 70-90% reduction | Medium-High | R² ~0.8-0.85 | Spectral data, collinear descriptors [26] |
Navigating Pitfalls and Enhancing Performance in QSAR Descriptor Selection
Troubleshooting Guides
Guide 1: Addressing Feature Selection Challenges in High-Dimensional Descriptor Spaces
Problem Statement: My QSAR model shows excellent training performance but fails to generalize to external test sets, likely due to overfitting from too many molecular descriptors relative to my compound count.
Diagnosis: This is a classic symptom of overfitting in high-dimensional, small-sample scenarios. The model is learning noise and spurious correlations instead of genuine structure-activity relationships.
Solution: Implement rigorous descriptor selection and model validation strategies.
Step-by-Step Resolution:
- Apply Feature Selection Methods: Use filter methods like SelectKBest or mutual information ranking to reduce descriptor space dimensionality before model building [57] [21].
- Incorporate Regularization: Utilize algorithms with built-in feature selection like LASSO (L1 regularization) or ensemble methods like Random Forests that provide feature importance metrics [41] [21].
- Validate with External Sets: Always hold out a completely external validation set before any model building to test generalizability [58].
- Apply Domain of Applicability: Use Williams plots or distance-based methods to identify when you're extrapolating beyond your model's reliable domain [57] [59].
Preventative Measures:
- Maintain a minimum compound-to-descriptor ratio of 5:1
- Use cross-validation strictly on training data only
- Apply multiple feature selection methods and compare results
Guide 2: Managing Small Sample Sizes in QSAR Modeling
Problem Statement: I have limited experimental data (less than 100 compounds) but need to build a predictive QSAR model for virtual screening.
Diagnosis: Small datasets are particularly vulnerable to overfitting and may not adequately represent the chemical space of interest.
Solution: Leverage transfer learning, data imputation, and specialized model architectures designed for small datasets.
Step-by-Step Resolution:
- Implement Multitask Learning: Train models on multiple related endpoints simultaneously, as demonstrated for CYP450 inhibition prediction where models trained on larger CYP isoform datasets improved performance for smaller CYP2B6 and CYP2C8 datasets [60].
- Use Data Imputation Techniques: For multitask settings, employ advanced imputation methods to handle missing values across related bioactivity datasets [60].
- Incorplement Quantum Mechanical Descriptors: Utilize QM descriptors like the QMex dataset, which have shown improved extrapolative performance for small-data molecular properties [59].
- Apply Interactive Linear Regression: For small datasets, use interpretable models like ILR with interaction terms between QM descriptors and structural categories [59].
Preventative Measures:
- Use simple, interpretable models when data is scarce
- Incorporate domain knowledge through meaningful descriptors
- Apply ensemble methods to reduce variance
Frequently Asked Questions
General Strategy Questions
Q: What is the most critical first step when building QSAR models with high-dimensional descriptors? A: The most critical step is rigorous descriptor selection before model building. Start with filter methods (SelectKBest, mutual information) to reduce dimensionality, followed by embedded methods like LASSO or Random Forests for further refinement [57] [21]. Never skip this step, especially with small sample sizes.
Q: How can I assess whether my model is overfitted? A: Look for these warning signs: (1) Large discrepancy between training and test set performance, (2) Poor performance on external validation sets, (3) Models that are overly complex relative to your data size, and (4) Feature importance rankings that highlight "bulk" properties rather than specific pharmacophoric features [5] [58].
Q: Are complex deep learning models better for small QSAR datasets? A: Generally no. Comprehensive benchmarks show that for small datasets (n < 500), simpler models like Interactive Linear Regression with QM descriptors often outperform deep learning models, which require large amounts of data to avoid overfitting [59]. Graph Neural Networks can be effective but require careful regularization and often benefit from transfer learning approaches [60].
Technical Implementation Questions
Q: What validation strategies are most appropriate for small datasets? A: With small datasets, use leave-one-out cross-validation (LOO-CV) or repeated k-fold cross-validation with multiple splits. Most importantly, always validate on a completely external test set that's never used in feature selection or parameter tuning [58] [61]. For virtual screening applications, focus on Positive Predictive Value (PPV) in the top predictions rather than balanced accuracy [58].
Q: How should I handle highly imbalanced datasets in classification QSAR? A: For virtual screening, avoid balancing your training set. Recent research shows that training on imbalanced datasets representative of real-world screening libraries produces higher positive predictive value in the top-ranked compounds—which is more important for experimental follow-up [58]. Focus on PPV rather than balanced accuracy for hit identification tasks.
Q: What molecular descriptors work best for small datasets? A: Quantum mechanical descriptors and 3D descriptors generally provide better extrapolative performance for small datasets compared to simple 2D descriptors [59] [62]. The QMex dataset and other QM descriptors capture electronic properties that are more mechanistically informative than simple topological descriptors [59].
Experimental Protocols & Methodologies
Protocol 1: Double Machine Learning for Deconfounding Molecular Descriptors
Purpose: To identify causal molecular descriptors rather than merely correlated ones in high-dimensional descriptor spaces [5].
Materials:
- Molecular dataset with biological activity measurements
- Computational chemistry software for descriptor calculation (RDKit, DRAGON)
- Python/R with machine learning libraries
Procedure:
- Calculate comprehensive molecular descriptor set (1000+ descriptors)
- Implement Double/Debiased Machine Learning (DML) framework:
- Stage 1: Regress each descriptor against all other descriptors using ML
- Stage 2: Regress biological activity against residuals from Stage 1
- Apply Benjamini-Hochberg procedure to control False Discovery Rate
- Identify descriptors with statistically significant causal effects
Validation:
- Compare against standard feature importance methods (Random Forest, LASSO)
- Test if "bulk" property proxies are correctly de-emphasized
- Verify identified descriptors make chemical sense
Protocol 2: Multitask Learning with Data Imputation for Small Datasets
Purpose: To improve predictive performance for small QSAR datasets by leveraging related bioactivity data through multitask learning [60].
Materials:
- Primary small dataset (target endpoint)
- Secondary larger datasets (related endpoints)
- Deep learning framework (PyTorch, TensorFlow)
- Graph neural network implementation
Procedure:
- Curate and align multiple related bioactivity datasets
- Implement missing data imputation for compounds without full activity profiles
- Design multitask neural network architecture:
- Shared hidden layers across all tasks
- Task-specific output layers
- Graph convolutional layers for molecular representation
- Train simultaneously on all available activities
- Fine-tune on primary target if needed
Validation:
- Compare against single-task baseline
- Use strict train/test splits per endpoint
- Evaluate extrapolative performance on novel scaffolds
Performance Comparison Tables
Table 1: Feature Selection Method Comparison for High-Dimensional Descriptor Spaces
| Method | Mechanism | Advantages | Limitations | Best Use Cases |
|---|---|---|---|---|
| SelectKBest | Selects K features highest statistical correlation with target [57] | Fast, interpretable, reduces dimensionality quickly | Univariate, may miss interactions | Initial descriptor screening |
| LASSO (L1) | Adds penalty term to regression that forces weak feature coefficients to zero [41] | Built-in feature selection, handles multicollinearity | May select only one from correlated features | Final model building with interpretation needs |
| Random Forest | Uses feature importance based on node impurity reduction [41] | Handles nonlinearities, robust to outliers | Biased toward high-cardinality features | Complex relationships, noisy data |
| DML Framework | Deconfounds features using double machine learning [5] | Identifies causal descriptors, reduces spurious correlations | Computationally intensive, complex implementation | Eliminating proxy variables, mechanistic insights |
Table 2: Model Performance with Small Sample Sizes (n < 500)
| Model Type | Extrapolation Performance* | Training Speed | Interpretability | Data Requirements |
|---|---|---|---|---|
| Interactive Linear Regression + QM | High [59] | Fast | High | Low (n ~ 100) |
| Random Forest | Medium [59] | Medium | Medium | Medium (n > 200) |
| Graph Neural Networks (Single Task) | Low [59] | Slow | Low | High (n > 1000) |
| Graph Neural Networks (Multitask) | Medium-High [60] | Slow | Low | Medium (with related data) |
*Extrapolation performance measured as maintenance of R² on external test sets outside training distribution [59]
Table 3: Key Computational Tools for Robust QSAR Modeling
| Tool/Resource | Function | Application Context | Key Features |
|---|---|---|---|
| RDKit | Molecular descriptor calculation [41] | General QSAR, descriptor generation | Open-source, comprehensive 2D/3D descriptors |
| QMex Dataset | Quantum mechanical descriptors [59] | Small dataset modeling, extrapolation | 150+ QM descriptors for organic molecules |
| SHAP Analysis | Model interpretation [57] | Feature importance analysis | Explains individual predictions, global interpretability |
| Double ML Framework | Causal descriptor identification [5] | High-dimensional descriptor spaces | Deconfounds correlated descriptors, hypothesis testing |
| Multitask GCN | Small dataset enhancement [60] | Limited data scenarios | Transfer learning across related endpoints |
Diagram: Strategic Approach Selection for QSAR Modeling
Troubleshooting Guides
Guide: Addressing Poor Model Performance and Overfitting
Problem: Your QSAR model performs well on training data but poorly on external test sets or new compounds, indicating potential overfitting and unreliable predictions.
Solution: Systematically evaluate and refine your molecular descriptors and modeling process.
| Step | Action | Key Checks & Quantitative Metrics |
|---|---|---|
| 1. Diagnose | Analyze performance disparity between training and validation sets. | Calculate ( R^2 ) (training) vs. ( Q^2 ) (cross-validation); a significant drop (e.g., ( R^2 > 0.9 ) while ( Q^2 < 0.5 )) suggests overfitting [63]. |
| 2. Review Descriptors | Check for irrelevant, redundant, or high-dimensional descriptors. | Assess descriptor contingency (target > 0.6) and correlation coefficients (e.g., Cramer's V > 0.2); remove those below thresholds [63]. Use feature selection to reduce dimensionality [27]. |
| 3. Validate & Test | Perform rigorous internal and external validation. | Use Leave-One-Out (LOO) cross-validation and calculate ( R^2 ), RMSE for fit and prediction [63]. Ensure predictions on a true external test set are reliable. |
| 4. Define Applicability Domain (AD) | Define the chemical space where the model makes reliable predictions. | Model predictions for a specific set of compounds are only considered reliable within this defined theoretical chemical space [55]. |
Guide: Selecting Mechanistically Interpretable Descriptors
Problem: The model identifies "bulk" properties (e.g., molecular weight) as highly predictive, but these are not actionable for chemists to design improved compounds, as they may be proxies for true causal features.
Solution: Implement a causal inference framework to move from correlational to causal QSAR.
| Step | Action | Key Checks & Quantitative Metrics |
|---|---|---|
| 1. Identify Confounders | Acknowledge that standard ML models can be misled by high-dimensional, correlated descriptors. | A "bulk" property may appear predictive but is merely correlated with the true, specific pharmacophore (e.g., a hydrogen bond donor) [5]. |
| 2. Deconfound Descriptors | Use statistical frameworks to estimate the unconfounded causal effect of each descriptor. | Apply Double/Debiased Machine Learning (DML) to treat all other descriptors as potential confounders [5]. |
| 3. Statistical Testing | Control for false discoveries and identify statistically significant causal links. | Apply the Benjamini-Hochberg procedure to the DML estimates to control the False Discovery Rate (FDR) and identify descriptors with a significant causal link to activity [5]. |
Frequently Asked Questions (FAQs)
Q1: What are the fundamental criteria for selecting high-quality molecular descriptors? Descriptors must comprehensively represent molecular properties, have a distinct chemical meaning, correlate with the biological activity, be computationally feasible, and be sensitive enough to capture subtle structural variations [27]. Balance between descriptor dimensions and computational cost is crucial to avoid the 'garbage in, garbage out' situation [27].
Q2: My dataset has incomplete property annotations (e.g., some ADMET properties missing for many compounds). How can I build a reliable model? Imperfectly annotated data is a common challenge. A unified multi-task learning framework like OmniMol can be effective. It formulates molecules and properties as a hypergraph, allowing the model to learn from all available molecule-property pairs simultaneously. This integrates correlations among different properties, enhancing the dataset's potential and leading to more robust predictions [64].
Q3: How can I ensure my model's predictions are explainable to guide chemists in synthesis? Modern frameworks like OmniMol are designed for explainability across three key relationships: among molecules, molecule-to-property, and among properties [64]. Furthermore, using 3D-QSAR models that provide favorable interaction maps (e.g., for H-bond acceptors/donors) in the binding site offers visual, actionable insights for hit optimization [65]. Prioritizing descriptors with a proven causal link to activity also enhances interpretability [5].
Q4: What is a standard experimental protocol for developing and validating a robust QSAR model? The following workflow outlines a standard protocol for QSAR model development and validation, integrating key steps from descriptor calculation to model application.
Q5: What are some common 2D descriptors and why are they used? Many QSAR models rely on a set of common, interpretable 2D descriptors that provide foundational information about a molecule's physicochemical character. The table below details several key examples.
| Descriptor Name | Brief Explanation | Function / Relevance |
|---|---|---|
| logP(o/w) | Log of the octanol/water partition coefficient. | Predicts lipophilicity, crucial for membrane permeability and absorption [63]. |
| TPSA | Topological Polar Surface Area. | Estimates a molecule's ability to engage in polar interactions, closely related to bioavailability and cellular permeability [63]. |
| a_acc | Number of hydrogen bond acceptor atoms. | Critical for estimating drug solubility and its interaction with biological targets [63]. |
| Molecular Weight | Mass of the molecule. | A fundamental property often correlated with bioavailability and other ADMET properties [63]. |
| Wiener Polarity Number | A topological index derived from the molecular graph. | Related to molecular branching and flexibility, which can influence binding [63]. |
The Scientist's Toolkit: Research Reagent Solutions
| Category / Tool | Specific Examples / Functions |
|---|---|
| Software & Platforms | MOE (for 2D descriptor calculation and QSAR modeling) [63]; OmniMol (unified multi-task framework for imperfect data) [64]; Orion, ROCS, EON (for 3D-QSAR featurization with shape and electrostatics) [65]. |
| Descriptor Types | 2D Descriptors: apol, bpol, a_heavy, logS [63]. 3D Descriptors: Molecular shape and electrostatic complementarity from tools like ROCS and EON [65]. |
| Advanced Modeling Frameworks | Double/Debiased Machine Learning (DML): A statistical framework for deconfounding molecular descriptors to identify causal features [5]. Hypergraph-based Models: To capture complex many-to-many relations between molecules and properties from imperfectly annotated data [64]. |
| Validation & Analysis | Genetic Function Approximation-Multiple Linear Regression (GFA-MLR): For developing robust QSAR models [66]. Benjamini-Hochberg Procedure: For controlling the False Discovery Rate (FDR) in high-dimensional hypothesis testing of descriptors [5]. |
FAQs and Troubleshooting Guides
This section addresses common challenges researchers face when defining the Applicability Domain (AD) of QSAR models.
FAQ 1: What does it mean if my new compound is flagged as an "outlier" or outside the AD? An outlier is a query compound that is structurally dissimilar to the compounds used to train your QSAR model. The principle of similarity states that predictions are reliable only for compounds similar to the training set [67]. Being outside the AD indicates that the model's prediction for this compound may be unreliable [68]. To troubleshoot:
- Action 1: Check the descriptors of the outlier compound against the range of each descriptor in your training set. The compound may have descriptor values outside the minimum or maximum values found in the training data [67].
- Action 2: Calculate the distance (e.g., Euclidean, Mahalanobis) from the outlier to the centroid of your training set. A distance greater than a pre-defined threshold (e.g., the maximum distance in the training set) confirms its outlier status [67].
FAQ 2: My model has good internal validation statistics, but poor predictive performance for new compounds. What is wrong? This often occurs when the new compounds fall outside your model's Applicability Domain. Good internal statistics only confirm the model's robustness for your training data; they do not guarantee predictions for structurally different molecules [69]. To resolve this:
- Action 1: Redefine your model's AD using a distance-based method like leverage or Mahalanobis distance. This will help you identify which new compounds are true extrapolations [67].
- Action 2: Ensure your original training set is diverse and representative of the chemical space you intend to predict. A narrow training set leads to a restricted AD [68].
FAQ 3: How can I quickly assess if my dataset is suitable for building a reliable QSAR model? Before model building, you can calculate the rivality and modelability indexes for your dataset. These indexes have a very low computational cost and do not require building a model [68].
- Rivality Index (RI): Assigns a value between -1 and +1 to each molecule. Molecules with high positive values are difficult to predict and likely outside the AD, while those with high negative values are easy to predict and lie within the AD [68].
- Interpretation: A dataset with many high positive RI values will be difficult to model reliably, signaling potential issues with the AD before you even begin [68].
FAQ 4: Which method should I choose to define the Applicability Domain of my model? The choice of method depends on your data and the trade-off between simplicity and comprehensiveness. The table below compares common approaches.
Table 1: Comparison of Key Applicability Domain (AD) Methods
| Method Category | Example | Key Principle | Advantages | Disadvantages |
|---|---|---|---|---|
| Range-Based | Bounding Box [67] | Defines a p-dimensional hyper-rectangle based on min/max descriptor values. | Simple and easy to implement. | Cannot identify empty regions or account for descriptor correlation. |
| Geometric | Convex Hull [67] | Defines the smallest convex area containing the entire training set. | Provides a defined geometric boundary. | Computationally complex for high-dimensional data; cannot identify internal empty regions. |
| Distance-Based | Leverage, Mahalanobis Distance [67] | Calculates the distance of a query compound from the training set's centroid. | Accounts for data distribution; Mahalanobis distance handles correlated descriptors. | Performance is highly dependent on the threshold setting. |
| Probability Density-Based | Probability Density Distribution [67] | Estimates the probability density of the training set in the descriptor space. | Accounts for the underlying data distribution. | Can be computationally intensive. |
| Advanced / Hybrid | Rivality Index (RI) [68] | Measures the capacity of each molecule to be correctly classified. | Low computational cost; no model building required; provides a local predictability measure. | Primarily demonstrated for classification models. |
Experimental Protocols for AD Assessment
This section provides detailed methodologies for key experiments related to defining and validating the Applicability Domain.
Protocol 1: Implementing a Distance-Based Applicability Domain using Leverage
1. Objective: To identify query compounds that are influential or outside the structural domain of the training set based on their leverage values. 2. Materials and Software: * A validated QSAR model (regression-based). * Training set descriptor matrix (X). * Query compound descriptor values. * Computational software (e.g., MATLAB, Python with NumPy). 3. Procedure: * Step 1: Center your training set descriptor matrix, X. * Step 2: Calculate the leverage matrix, H, using the formula: H = X(XᵀX)⁻¹Xᵀ [67] * Step 3: The diagonal values of the H matrix are the leverage values for each compound. Calculate the warning leverage (h), typically defined as: h = 3p/N where p is the number of model descriptors plus one, and N is the number of training compounds [67]. * Step 4: For a new query compound, calculate its leverage (hᵢ). If hᵢ > h*, the compound is considered influential and outside the AD, and its prediction should be treated as unreliable [67].
Protocol 2: Calculating the Rivality Index (RI) for a Classification Dataset
1. Objective: To assess the modelability of a dataset and identify compounds difficult to predict prior to model building. 2. Materials and Software: * A dataset with molecular structures and a categorical biological activity. * Software capable of calculating molecular descriptors and the RI index. 3. Procedure: * Step 1: Compute molecular descriptors for all compounds in the dataset. * Step 2: For each molecule i in the dataset, identify its nearest neighbor that belongs to the same class and its nearest neighbor that belongs to the opposite class [68]. * Step 3: The Rivality Index for molecule i is calculated based on the relative distances to these two neighbors. The index value falls in the range [-1, +1] [68]. * Step 4: Interpret the results: * RI ≈ -1: The molecule is easy to predict and lies firmly within the AD. * RI ≈ +1: The molecule is difficult to predict (an outlier) and resides outside the AD [68]. * A dataset with many high positive RI values has low modelability and will likely produce a model with a narrow AD.
Protocol 3: Defining the AD using the PCA Bounding Box Method
1. Objective: To define the AD in a way that accounts for correlation between descriptors. 2. Materials and Software: * Training set descriptor matrix. * Software for Principal Component Analysis (PCA). 3. Procedure: * Step 1: Perform PCA on the descriptor matrix of the training set. * Step 2: Select the number of significant principal components (PCs) that capture most of the variance in the data. * Step 3: For each significant PC, record the minimum and maximum scores from the training set projections. This defines a hyper-rectangle in the PC space [67]. * Step 4: For a new query compound, project its descriptors onto the same PCs. If the compound's score on any PC falls outside the min-max range of the training set for that PC, it is considered outside the AD [67].
AD Assessment Workflow
The diagram below outlines a logical workflow for assessing whether a new compound falls within your model's Applicability Domain.
Research Reagent Solutions
Table 2: Essential Computational Tools for QSAR and Applicability Domain
| Item / Software | Function in QSAR/AD Studies |
|---|---|
| Molecular Descriptors (e.g., from Mordred Python package) [34] | Provide a quantitative, numerical representation of molecular structures, which are the fundamental inputs for building QSAR models and defining the Applicability Domain. |
| QSAR Modeling Software (e.g., with SVM, Random Forest) [68] | Algorithms used to build the mathematical relationship between molecular descriptors and biological activity. The choice of algorithm can influence the model's performance and AD. |
| Validation Tools (e.g., Double Cross-Validation, Y-Scrambling) [69] [70] | Techniques and software used to assess the robustness and predictive power of a QSAR model, which is a prerequisite for properly defining its AD. |
| Applicability Domain Methods (e.g., Leverage, RI, PCA Bounding Box) [68] [67] | Specific algorithms and scripts used to characterize the interpolation space of a model and identify outliers, ensuring reliable predictions. |
Frequently Asked Questions
Q1: How can I tell if my QSAR data has a nonlinear relationship that requires a specialized modeling approach? A strong indicator is when simple linear models, like Multiple Linear Regression (MLR), show poor performance (low R² and high prediction error) on your training data, but more complex models demonstrate significantly better results [71] [72]. For instance, in a study predicting hERG channel inhibition, a Gradient Boosting model substantially outperformed a linear model, indicating underlying nonlinearity or complex descriptor interactions that the linear model could not capture [72]. A noticeable difference between the performance on the training set and the validation/test set can also suggest the model is struggling to generalize, which may be due to unaccounted-for nonlinearity.
Q2: My dataset has a large number of descriptors. What is a robust method to select the most relevant ones for a nonlinear model? For high-dimensional descriptor spaces, Recursive Feature Elimination (RFE) is a powerful, supervised technique. Unlike simple filtering based on variance or correlation, RFE iteratively builds a model (like a Gradient Boosting Machine) and removes the least important descriptors based on their impact on model performance [72] [41]. This ensures that the final set of descriptors is predictive in the context of the full model and the target property. This method is particularly effective for avoiding overfitting while retaining informative features [73].
Q3: What should I do if my molecular descriptors are highly correlated with each other (multicollinearity) before building a nonlinear model? While some advanced nonlinear models like Gradient Boosting Machines are inherently robust to descriptor correlation, it is still good practice to address it [72]. You can:
- Generate a correlation matrix to identify and manually remove one descriptor from each pair of highly correlated descriptors.
- Use feature selection methods like RFE, which naturally down-weights redundant descriptors during the iterative training process [72].
- Apply regularization techniques embedded in some algorithms that penalize complex models, thus indirectly handling redundancy [41].
Q4: Are there specific types of molecular descriptors that are better suited for capturing nonlinear relationships? Yes, complex relationships often require descriptors that encode richer information about the molecule. While traditional 2D descriptors (e.g., logP, topological indices) can be used, 3D field descriptors (like Cresset's XED fields) that capture a molecule's shape and electrostatic character as a protein "sees" it can be highly effective [72]. Furthermore, modern AI-derived "deep descriptors" from Graph Neural Networks (GNNs) or other deep learning models automatically learn complex, hierarchical features directly from molecular structure, making them exceptionally powerful for modeling nonlinear structure-activity relationships [41].
Q5: What is a key advantage of using a nonlinear method like Gene Expression Programming (GEP) over a linear model? Nonlinear methods like GEP automatically generate complex, symbolic relationships between your descriptors and the biological activity without relying on pre-defined linear equations. In a study on osteosarcoma, a GEP model (R²=0.839) showed much greater consistency with experimental values than a linear heuristic model (R²=0.603), demonstrating its superior ability to capture the underlying complex activity landscape [71].
Troubleshooting Guides
Problem: Poor Model Performance and Inability to Capture Complex Patterns
Symptoms:
- Low R² values on both training and test sets.
- A simple linear model performs poorly, but a more complex model shows significantly better results [72].
- Visual inspection of predicted vs. actual values shows a curved pattern of residuals, not a random scatter.
Diagnosis: The relationship between your molecular descriptors and the target endpoint is likely nonlinear and cannot be adequately captured by a linear model [71] [41].
Solution: Adopt a nonlinear machine learning model and ensure your descriptors are capable of encoding complex information.
Recommended Steps:
- Switch to a Nonlinear Algorithm: Implement a model designed for nonlinearity, such as:
- Gradient Boosting Machines (GBM): Highly effective and robust to correlated descriptors [72].
- Random Forests (RF): Good for handling noisy data and provides built-in feature importance [41].
- Gene Expression Programming (GEP): Excellent for automatically discovering complex symbolic relationships [71].
- Graph Neural Networks (GNNs): For an advanced approach that uses the molecular graph directly [41].
- Expand Your Descriptor Set: Move beyond basic 1D/2D descriptors. Calculate and incorporate 3D field descriptors or use AI-generated deep descriptors for a more comprehensive molecular representation [72] [41].
- Validate Rigorously: Use a strict train-test split and k-fold cross-validation to ensure your nonlinear model generalizes well and is not overfitting [72] [73].
Problem: Model Overfitting Due to High-Dimensional Descriptor Space
Symptoms:
- The model performs excellently on the training data but poorly on the test data or new compounds.
- A large delta (difference) between training and testing R² and RMSE values.
Diagnosis: The model has become too complex and has learned the noise in the training data, often due to too many irrelevant or redundant descriptors [72] [73].
Solution: Implement a robust feature selection strategy to reduce dimensionality.
Recommended Steps:
- Initial Descriptor Filtering:
- Apply Supervised Feature Selection:
- Use Regularization:
- Employ algorithms that incorporate regularization (e.g., LASSO) to penalize model complexity and drive the coefficients of less important descriptors toward zero [41].
Protocol: A Workflow for Building a Robust Nonlinear QSAR Model
This protocol outlines a systematic approach for developing a QSAR model when nonlinear relationships are suspected, integrating best practices from recent literature [72] [73].
Table 1: Key Diagnostic Indicators for Linear vs. Nonlinear QSAR Models
This table helps diagnose whether your data requires a nonlinear modeling approach based on a case study of hERG inhibition prediction [72].
| Diagnostic Metric | Linear Regression Model Performance | Gradient Boosting Model Performance | Interpretation |
|---|---|---|---|
| R² (Test Set) | Low | > 0.5 | Nonlinear model captures complex patterns linear model misses. |
| Root Mean Squared Error (RMSE) | High | Low | Nonlinear model predictions are closer to experimental values. |
| R² Delta (Train vs. Test) | Small | ~0.04 | Low delta in a complex model suggests good generalization and less overfitting. |
| RMSE Delta (Train vs. Test) | Small | ~6.6% | Consistent performance across training and test sets indicates a robust model. |
Table 2: Suitability of Different Descriptor Types for Nonlinear QSAR
This table summarizes various molecular descriptors and their applicability for modeling complex, nonlinear relationships [72] [41].
| Descriptor Class | Examples | Advantages | Considerations for Nonlinear Models |
|---|---|---|---|
| Physicochemical (1D) | Molecular weight, logP, H-bond donors/acceptors | Fast to compute; easily interpretable. | May be insufficient to capture complex activity on their own. |
| Topological & 2D Fingerprints | Kier-Hall indices, Morgan fingerprints | Encode molecular structure and substructures; no 3D conformation needed. | Excellent for nonlinear models; provide a rich feature set for pattern recognition. |
| 3D Field Descriptors | Cresset XED (electrostatic, shape) | Represents how a protein "sees" the ligand; high information content. | Requires a bioactive conformation; powerful for capturing subtle steric/electronic effects. |
| Quantum Chemical | HOMO/LUMO energies, dipole moment, electrostatic potential | Describe electronic properties crucial for reactivity and binding. | Computationally intensive; can be highly informative for specific target classes. |
| AI-Derived (Deep Descriptors) | Graph Neural Network (GNN) embeddings, SMILES-based latent representations | Automatically learned; capture hierarchical features without manual engineering. | State-of-the-art; requires larger datasets and more computational resources. |
The Scientist's Toolkit: Essential Research Reagents & Software
Table 3: Key Software and Tools for Nonlinear QSAR Modeling
| Tool Name | Function | Application in Descriptor Selection |
|---|---|---|
| CODESSA | Calculates a wide range of molecular descriptors (topological, electrostatic, quantum mechanical) [71]. | Used for comprehensive descriptor generation prior to model building. |
| RDKit | Open-source cheminformatics toolkit. | Calculates 2D and 3D descriptors and fingerprints; often integrated into other platforms like Flare [72]. |
| Flare (Cresset) | Commercial software for 3D-QSAR and molecular modeling. | Provides 3D field descriptors and built-in Gradient Boosting models for robust nonlinear QSAR [72]. |
| KNIME | Open-source data analytics platform with extensive cheminformatics extensions. | Facilitates the creation of automated QSAR workflows, including feature selection and model validation [73]. |
| Python (scikit-learn, XGBoost) | Programming language with powerful machine learning libraries. | Offers full control for implementing custom feature selection (RFE) and advanced nonlinear algorithms [72] [41]. |
Troubleshooting Guides
Algorithm Implementation Issues
Q: The dynamic importance adjustment in our CPANN model is not converging. The model performance is unstable during training. What could be the cause?
A: Non-convergence often stems from issues with the dynamic scaling factor m(t, i, j, k) in the weight update equation. Adhere to the following protocol to ensure stability [25]:
- Verify the Scaling Factor Calculation: Ensure the term
m(t, i, j, k)is computed correctly using the equation:m(t, i, j, k) = [1 − (1 − p(t)) ∙ ABS[scaled(o(k)) − scaled(w(i, j, k))]] ∙ [1 − (1 − p(t)) ∙ ABS[scaled(o(target)) − scaled(w(i, j, target))]]Confirm thatp(t)decreases linearly from 1 to 0 during training [25]. - Inspect Input Data Scaling: The algorithm uses range-scaled values for the object's descriptor
scaled(o(k)), neuron weightscaled(w(i, j, k)), and target propertyscaled(w(i, j, target)). Improper scaling of input data can lead to numerical instability and failed convergence [25] [3]. - Adjust Training Parameters: The learning coefficient
η(t)should linearly decrease from a predefined maximum to a minimum value. An excessively high initial learning rate can cause oscillations, while a rate that is too low can stall convergence [25].
Q: Our model's interpretability is poor despite using dynamic descriptor importance. How can we identify which molecular features the model deems most critical?
A: To enhance interpretability, integrate post-hoc explanation techniques with your dynamic model [25] [41] [74]:
- Employ SHAP or LIME: Use SHapley Additive exPlanations (SHAP) or Local Interpretable Model-agnostic Explanations (LIME) on the trained model. These methods can help identify which descriptors were most influential for predictions, even in complex non-linear models like neural networks [41] [74].
- Analyze Neuron Weights: In CPANN, directly examine the final importance values of molecular descriptors on the excited neurons for different endpoint classes. This can reveal key molecular features responsible for classifying molecules into specific categories [25].
- Validate with Known Structural Alerts: Cross-reference the high-importance descriptors identified by the model with known structural alerts or mechanistic information from the literature. For example, a model might correctly assign high importance to descriptors like
nRNNOx(number of N-nitroso groups) for carcinogenicity, which is a known structural alert [25].
Data and Feature Management
Q: What is the best strategy for selecting an initial set of molecular descriptors before applying dynamic importance adjustment?
A A hybrid approach that combines feature selection with feature learning often yields the best results [75]:
- Leverage Feature Selection Tools: Use software like DELPHOS or DRAGON to perform an initial feature selection from a large pool of computed descriptors (e.g., 0D, 1D, and 2D descriptors). This reduces dimensionality and computational load [75].
- Incorporate Feature Learning: Complement the selected descriptors with features learned by algorithms like CODES-TSAR, which generates numerical descriptors directly from the SMILES representation of a molecule. This can capture complementary information not present in traditional descriptors [75].
- Utilize Embedded Methods: For classical models, apply embedded feature selection methods like LASSO regression, which performs feature selection as part of the model training process. This is an efficient way to identify the most relevant variables [41] [74].
Q: How should we handle missing values in our molecular descriptor dataset to prevent errors in dynamic adjustment algorithms?
A Robust data preprocessing is essential [3]:
- Imputation Techniques: For datasets with a low fraction of missing data, employ imputation methods such as k-nearest neighbors (k-NN) or matrix factorization to estimate and fill in the missing values [3].
- Removal of Compounds: If the number of compounds with missing data is small and the dataset is sufficiently large, consider removing those compounds entirely to maintain data integrity [3].
- Data Cleaning: Prior to modeling, rigorously clean and preprocess the dataset. This includes standardizing chemical structures (e.g., removing salts, normalizing tautomers) and converting all biological activities to a common unit and scale [3].
Software and Technical Setup
Q: The QSAR Toolbox application window fails to open after the splash screen appears. How can I resolve this?
A This is a known issue, often related to the .NET framework or regional settings [76] [77]:
- Follow BadImageFormatException Fix: This problem can manifest as a
System.BadImageFormatException. The official support site provides a dedicated document, "Toolbox Client starts but hides after the splash screen (BadImage)," which contains step-by-step resolution instructions [76]. - Check Regional Settings: On some operating systems with a display language different from English, the Toolbox may deadlock. Applying the official patch for the
DatabaseDeployermay resolve this issue [76].
Q: The QSAR Toolbox Server cannot connect to the PostgreSQL database when they are deployed on separate machines. What is the solution?
A This is a configuration issue with the PostgreSQL server's access controls [76]:
- On the machine hosting the PostgreSQL database, navigate to the PostgreSQL data folder (e.g.,
C:\Program Files\PostgreSQL\9.6\data) and open thepg_hba.conffile [76]. - Add a line to the bottom of the file to allow connection from the Toolbox Server's host. Replace
<ToolboxServerHost>with the IP address or hostname of the QSAR Toolbox Server machine [76]:host all qsartoolbox <ToolboxServerHost> md5 - Save the file and restart the PostgreSQL service [76].
- Restart the QSAR Toolbox Server application or service [76].
Experimental Protocols for Key Methodologies
Protocol: Implementing Dynamic Descriptor Importance in CPANN
This protocol details the procedure for implementing the novel dynamic descriptor importance adjustment in a Counter-Propagation Artificial Neural Network (CPANN) as described in the foundational research [25].
1. Model Setup and Initialization
- Network Architecture: Define a 2D Kohonen layer with dimensions
Nx × Ny. Each neuron hasNdescweights corresponding to the number of molecular descriptors. The output (Grossberg) layer has the same dimensions, with each neuron havingNtarweights (one for the target property in classification tasks) [25]. - Initialization: Initialize weights in the Kohonen and Grossberg layers to small random values [25].
- Parameters: Set the initial learning coefficient
η(t)and the triangular neighborhood functionh(i, j, t). Initializep(t)to 1, which will linearly decrease to 0 during training [25].
2. Dynamic Training Cycle
For each iteration t and each training molecule [25]:
- Find Winning Neuron: Calculate the Euclidean distance between the input molecule's descriptor vector and the weight vector of every neuron in the Kohonen layer. The neuron with the smallest distance is the central (winning) neuron [25].
- Calculate Dynamic Scaling Factor: For each descriptor
kand each neuron within the neighborhood, compute the dynamic scaling factorm(t, i, j, k)using the provided equation, which incorporates the differences between the scaled input values and neuron weights for both the descriptor and the target property [25]. - Update Kohonen Layer Weights: Adjust the weights of all neurons within the neighborhood of the central neuron using the formula:
w(t, i, j, k) = w(t − 1, i, j, k) + m(t, i, j, k) ∙ η(t) ∙ h(i, j, t) ∙ (o(k) − w(t − 1, i, j, k))The extent of the adjustment is governed by the learning rate, neighborhood function, and the dynamic scaling factor [25]. - Update Grossberg Layer Weights: Project the central neuron's position to the Grossberg layer. Adjust the weights in this layer using the same equation as above, but replace the molecular descriptor value
o(k)with the target property value of the input molecule [25]. - Update Parameters: Decrease the learning rate
η(t)and the parameterp(t)linearly according to the training schedule [25].
3. Model Validation
- Use internal validation (e.g., k-fold cross-validation) on the training set to optimize parameters [25] [3].
- Evaluate the final model's performance on a held-out external test set to assess its predictive power and generalizability [25] [3].
Protocol: Hybrid Feature Selection and Learning for QSAR
This protocol outlines a hybrid strategy to generate optimal molecular descriptor sets by combining feature selection and feature learning [75].
1. Data Preparation
- Compile a dataset of chemical structures (e.g., as SMILES strings) and their associated biological activities. Ensure the dataset is curated, cleaned, and standardized [3].
- Split the dataset into training, validation, and external test sets using a method like Kennard-Stone to ensure representative chemical space coverage [75] [3].
2. Feature Selection Pathway
- Descriptor Calculation: Use software like DRAGON or PaDEL-Descriptor to compute a comprehensive set of 0D, 1D, and 2D molecular descriptors for all compounds in the dataset [75] [3].
- Feature Selection: Apply a feature selection tool like DELPHOS to the calculated descriptors. DELPHOS splits the selection task into two phases to manage computational effort, identifying a reduced subset of descriptors (
D − D MD Sets) well-correlated with the target property [75].
3. Feature Learning Pathway
- Descriptor Generation: Process the SMILES strings of all compounds using the CODES tool. CODES creates a dynamic matrix from the molecular structure [75].
- Dimensionality Reduction: Input the matrix generated by CODES into the TSAR software. TSAR performs dimensionality reduction to compute a set of novel molecular descriptors (
C − T MD Sets) for each compound [75].
4. Hybridization and Modeling
- Descriptor Fusion: Create a combined descriptor set (
Both MD Sets) by merging the descriptors from the feature selection (D − D MD Sets) and feature learning (C − T MD Sets) pathways [75]. - Model Building: Use the combined descriptor set to build QSAR models with various machine learning algorithms (e.g., Random Forest, Support Vector Machines, Neural Networks) implemented in platforms like WEKA [75].
- Performance Assessment: Validate and compare the models built from the feature selection set, the feature learning set, and the combined set using the external test set. The combined set often achieves higher accuracy due to complementary information [75].
Table 1: Performance Comparison of Descriptor Set Strategies on Benchmark Datasets
This table summarizes the findings from a comparative study on hybrid feature strategies, showing how combining descriptor sets can improve model performance [75].
| Dataset (Target Property) | Best Model Type | Sampling Size | Strategy for MD Set | Key Performance Metric (Result) |
|---|---|---|---|---|
| Blood-Brain Barrier (BBB) | Regression | 75/25 | Both MD Sets | Correlation Coefficient (CC): 0.91 |
| Blood-Brain Barrier (BBB) | Classification | 66/34 | Both MD Sets | % Correctly Classified (%CC): 94.1% |
| Human Intestinal Absorption (HIA) | Regression | 75/25 | C − T MD Sets | Correlation Coefficient (CC): 0.89 |
| Human Intestinal Absorption (HIA) | Classification | 66/34 | D − D MD Sets | % Correctly Classified (%CC): 92.3% |
| Enantiomeric Excess (EE) | Regression | 50/50 | D − D MD Sets | Correlation Coefficient (CC): 0.85 |
| Enantiomeric Excess (EE) | Classification | 75/25 | C − T MD Sets | % Correctly Classified (%CC): 89.7% |
Table 2: Essential Research Reagent Solutions for Dynamic QSAR Modeling
A list of key software tools and databases essential for implementing dynamic descriptor adjustment and related QSAR methodologies.
| Item Name | Type | Primary Function / Application |
|---|---|---|
| DRAGON | Software | Calculates thousands of molecular descriptors (0D-3D) for a given set of compounds. Used for the initial feature pool in feature selection strategies [75] [41]. |
| PaDEL-Descriptor | Software | An open-source software for calculating molecular descriptors and fingerprint patterns. Useful for generating a wide array of 2D descriptors [75] [3]. |
| DELPHOS | Software | A feature selection method designed specifically for QSAR modeling. It efficiently identifies a reduced set of relevant molecular descriptors from a large initial pool [75]. |
| CODES-TSAR | Software | A feature learning method that generates numerical descriptors directly from SMILES codes, avoiding pre-defined molecular descriptors. Captures non-linear relationships [75]. |
| RDKit | Software Toolkit | An open-source cheminformatics toolkit that can be used for descriptor calculation, fingerprint generation, and molecular operations within custom Python scripts [41] [3]. |
| QSAR Toolbox | Software Platform | A regulatory tool that provides a workflow for profiling chemicals, defining categories, and filling data gaps via read-across. Aids in mechanistic interpretation [76] [8]. |
| LiverTox Database | Database | A curated database of drug-induced liver injury. Provides a source of hepatotoxicity data for building and validating classification models [25]. |
Workflow and System Diagrams
CPANN-v2 Dynamic Importance Training
Hybrid Feature Strategy Workflow
Ensuring Predictive Power: A Rigorous Framework for QSAR Model and Descriptor Validation
In Quantitative Structure-Activity Relationship (QSAR) modeling, a high R² value on training data can create a false sense of security. While the coefficient of determination (R²) measures how well a model fits the data it was trained on, it provides no guarantee of predictive performance on new, unseen chemical compounds [78]. This is especially critical in molecular design for drug discovery, where models must generalize beyond the training set to reliably predict the activity of novel compounds [27] [74].
External model validation represents the most rigorous assessment of a QSAR model's utility in real-world applications [79]. By testing models against completely independent datasets that were not used during model building or parameter tuning, researchers can obtain an honest estimate of predictive power [78] [79]. For QSAR models used in regulatory decisions or pharmaceutical development, moving beyond R² to comprehensive validation criteria is not merely academic—it is essential for ensuring chemical safety and reducing costly late-stage failures in drug development [54] [74].
Key Statistical Criteria for External Validation
When performing external validation, several statistical measures provide a more complete picture of model performance than R² alone. The following criteria are particularly valuable for assessing predictive ability in QSAR models.
Table 1: Key Statistical Metrics for External QSAR Model Validation
| Metric | Calculation | Interpretation | Optimal Value |
|---|---|---|---|
| Predicted R² | R²predict = 1 - (SSE/SST) [79] | Proportion of variance in external set explained by model | Close to 1 |
| Concordance Correlation Coefficient (CCC) | CCC = (2 × sxy) / (sx² + sy² + (x̄ - ȳ)²) | Measures agreement between observed and predicted values | Close to 1 |
| rm² Metrics | rm² = r² × (1 - √(r² - r₀²)) [27] | Combines correlation and slope considerations | > 0.5 |
| Q²F1, Q²F2, Q²F3 | Variations considering mean, variance differences | Different perspectives on predictive performance | > 0.5 |
Golbraikh-Tropsha Acceptance Criteria
The Golbraikh-Tropsha criteria represent a comprehensive framework for establishing model validity. A QSAR model is considered predictive if it satisfies ALL the following conditions:
- Cross-validated R² (Q²) > 0.5 - This ensures reasonable internal predictive ability [27]
- External R² (r²ext) > 0.5 - The model maintains performance on the external test set [27]
- Slope (k or k') of regression lines through origin between observed vs. predicted values should satisfy 0.85 < k < 1.15 [27]
- Coefficient of determination (r²₀ or r'²₀) between observed vs. predicted values through origin should be close to r²ext [27]
These criteria collectively ensure that the model demonstrates both correlation and accuracy in its predictions, going beyond what R² alone can reveal.
Concordance Correlation Coefficient (CCC)
The CCC evaluates both precision and accuracy by measuring how far observations deviate from the line of perfect concordance (the 45° line through the origin). Unlike Pearson's r, which only measures correlation, CCC also accounts for systematic bias in predictions [27]. For QSAR models, CCC values above 0.85 are generally considered excellent, while values below 0.65 indicate poor predictive ability.
The rm² Metrics and Their Interpretation
The rm² metrics (rm²(overall), rm²(delta), and rm²(average)) provide nuanced insights into model performance:
- rm²(overall): Combines information about both the correlation coefficient and the slope of the regression line
- rm²(delta): The difference between rm²(overall) and rm²(average), with values > 0.2 suggesting significant prediction bias
- Thresholds: rm²(overall) > 0.5 and rm²(delta) < 0.2 indicate acceptable predictive performance [27]
Experimental Protocol for Comprehensive External Validation
Implementing a rigorous external validation protocol requires careful planning and execution. The following workflow provides a systematic approach applicable to QSAR modeling in cheminformatics and drug discovery.
Diagram 1: External validation workflow for QSAR models
Data Splitting Methodology
Proper dataset division is fundamental to meaningful external validation. The external test set must remain completely untouched during model development and parameter optimization.
Kennard-Stone Algorithm: This method ensures that the external test set adequately represents the chemical space covered by the training set. It selects test compounds that span the entire descriptor space, preventing extrapolation beyond the model's applicability domain [3].
Y-Randomization Test: To confirm model robustness rather than chance correlation, the Y-randomization test shuffles biological activity values while keeping descriptor values intact. A valid model should perform poorly on randomized data, with R² and Q² values significantly lower than for the original data [27].
Minimum Sample Size Requirements
For statistically meaningful external validation:
- External test set: Minimum of 15-20 compounds to obtain reliable performance estimates [79]
- Training set: Sufficiently large to capture structure-activity relationships, typically 80-90% of available data
- Chemical diversity: Both sets should represent similar chemical space distributions to avoid extrapolation
The Scientist's Toolkit: Essential Research Reagents
Table 2: Essential Tools for QSAR Model Development and Validation
| Tool/Category | Specific Examples | Primary Function | Application in Validation |
|---|---|---|---|
| Descriptor Calculation | PaDEL-Descriptor, Dragon, RDKit, Mordred [3] | Generate molecular descriptors from chemical structures | Ensure consistent descriptor calculation across training and test sets |
| Modeling Algorithms | Multiple Linear Regression (MLR), Partial Least Squares (PLS), Random Forest, Support Vector Machines (SVM) [3] [74] | Build QSAR models using various mathematical approaches | Compare model performance across algorithms |
| Validation Software | QSARINS, Build QSAR [74] | Implement validation protocols and calculate metrics | Automate calculation of Golbraikh-Tropsha criteria, rm² metrics |
| Chemical Databases | PubChem, ChEMBL [27] | Source of structural and activity data | Provide external test sets for validation |
| Visualization Tools | SHAP, LIME [74] | Interpret model predictions and descriptor contributions | Understand model behavior on external compounds |
Troubleshooting Guide: Common Validation Issues and Solutions
FAQ 1: Why does my model have high R² but fails external validation?
Problem: The model shows excellent performance on training data (R² > 0.9) but performs poorly on the external test set (R²ext < 0.5).
Root Causes:
- Overfitting: The model has learned noise rather than true structure-activity relationships, often due to too many descriptors relative to training compounds [78]
- Inadequate applicability domain: External compounds differ structurally from training set molecules [27]
- Data leakage: Information from the test set inadvertently influenced model development [3]
Solutions:
- Apply stricter feature selection to reduce descriptor count [3]
- Use applicability domain assessment to identify where predictions are reliable [27]
- Verify dataset splitting was performed before ANY model building steps [79]
- Try simpler models (MLR, PLS) before complex machine learning approaches [74]
FAQ 2: How can I determine my model's applicability domain?
Problem: Uncertainty about which chemical structures the model can reliably predict.
Assessment Methods:
- Leverage approach: Calculate Hat matrix for training set, establish critical leverage threshold [27]
- Distance-based methods: Use Euclidean or Mahalanobis distance in descriptor space to identify outliers [27]
- PCA-based approach: Project compounds into principal component space, define boundaries based on training set distribution [74]
Implementation:
Diagram 2: Applicability domain (AD) assessment workflow
FAQ 3: What should I do when my model fails the Golbraikh-Tropsha criteria?
Problem: The model fails one or more of the Golbraikh-Tropsha acceptance criteria during external validation.
Diagnostic Steps:
- Check slope (k) criteria failure: This indicates systematic bias in predictions
- Solution: Examine descriptor distributions between training and test sets
- Consider data transformation or weighted regression approaches [27]
Address poor r²ext values: The model lacks general predictive ability
- Solution: Expand training set chemical diversity
- Re-evaluate descriptor selection—may be missing critical molecular features [3]
Improve r²₀ performance: Predictions show inconsistent error patterns
- Solution: Investigate potential activity cliffs or nonlinear relationships
- Consider nonlinear modeling approaches if data supports it [74]
FAQ 4: How do I choose between linear and nonlinear models for better external prediction?
Problem: Uncertainty about whether linear or nonlinear approaches will yield better externally predictive models.
Decision Framework:
- Use linear models (MLR, PLS) when:
- Consider nonlinear models (SVM, Random Forest, Neural Networks) when:
Validation Approach: Use exactly the same external test set to compare linear and nonlinear models, evaluating using multiple metrics beyond R² [78].
FAQ 5: What are the minimum reporting requirements for external validation in publications?
Problem: Uncertainty about what validation evidence should be reported to demonstrate model credibility.
Essential Reporting Elements:
- Dataset characteristics: Size and diversity of both training and external test sets [27]
- Data splitting method: Rationale and methodology for creating external test set [79]
- Complete validation metrics: R²train, Q², R²ext, CCC, rm² values, and Golbraikh-Tropsha criteria results [27]
- Applicability domain definition: Method used and percentage of external compounds within domain [27]
- Comparison to existing models: Performance relative to previously published approaches [74]
Comprehensive external validation moves QSAR modeling from mathematical exercise to practical tool for drug discovery [74]. By implementing the Golbraikh-Tropsha criteria, CCC, and rm² metrics alongside traditional measures, researchers can develop models with proven predictive ability [27]. This rigorous approach is especially critical when selecting molecular descriptors for QSAR research, as it reveals which descriptor sets genuinely capture structure-activity relationships rather than merely fitting training data [3].
As AI and machine learning transform QSAR modeling [74], robust external validation becomes even more crucial for distinguishing true predictive advances from sophisticated overfitting. By adopting these comprehensive validation practices, researchers can build QSAR models that reliably accelerate molecular design and drug development while minimizing costly experimental failures.
Troubleshooting Guide: QSAR Validation Techniques
This guide addresses common challenges researchers face when validating Quantitative Structure-Activity Relationship (QSAR) models, providing targeted solutions to ensure reliable assessment of model predictive power.
FAQ 1: Why does my external validation performance vary dramatically each time I run it with a different random split?
- Problem: High variability in external validation metrics (like R² for the test set) across different data splits.
- Cause: This instability is particularly pronounced in datasets with a small number of samples but a large number of molecular descriptors (a common scenario in QSAR) [80]. The variation arises because a single, random train-test split may not be representative of the entire, limited dataset. A different split can lead to a test set that is either easier or harder to predict, significantly impacting the performance metrics [80] [81].
- Solution:
- For small datasets (n < 100 compounds), prefer Leave-One-Out (LOO) cross-validation. A comparative study found LOO to have the most stable and best overall performance for high-dimensional, small-sample data [80].
- If using external validation, perform multiple data splits (e.g., 10-20 random splits) and report the average and standard deviation of the performance metrics. This provides a more realistic view of model stability [80].
- Ensure your dataset is as large and diverse as possible before splitting.
FAQ 2: A high coefficient of determination (r²) for my test set suggests a good model, but the predictions seem poor. Why is this misleading?
- Problem: A high r² value during external validation does not guarantee a model with reliable predictive power [81].
- Cause: The r² metric alone is an insufficient indicator of model validity. It can be inflated by outliers or might not reflect the model's accuracy across the entire range of activity values. Other statistical characteristics must be considered [81].
- Solution: Do not rely on a single metric. Adopt a multi-faceted validation approach that includes several of the following for the external test set [81]:
- Calculate metrics like
r₀²andr'₀²(coefficients of determination for regression through the origin) and check if they are close to each other. - Examine the absolute error distribution and its standard deviation.
- Ensure that the mean and variance of the predicted activities are consistent with those of the experimental activities.
- Calculate metrics like
FAQ 3: When building a model with many descriptors, how do I avoid overfitting and ensure it will generalize to new compounds?
- Problem: The model performs excellently on the training data but fails to predict new, unseen compounds accurately.
- Cause: Overfitting occurs when a model learns the noise in the training data rather than the underlying structure-activity relationship. This is a major risk when the number of molecular descriptors (
p) is much larger than the number of compounds (n) [80] [82]. - Solution:
- Apply Feature Selection: Use feature selection methods (e.g., genetic algorithms, LASSO regression) to identify and use only the most relevant molecular descriptors [83] [82]. LASSO is particularly useful as it performs variable selection and modeling simultaneously [80].
- Use Rigorous Internal Validation: Employ k-fold cross-validation (e.g., 5-fold or 10-fold) during the model training phase to tune parameters and select features. This helps ensure the model captures generalizable patterns [84].
- Define an Applicability Domain: Finally, test the finalized model on a completely held-out external test set to obtain an unbiased estimate of its predictive performance [3].
Comparative Performance of Validation Techniques
The table below summarizes a quantitative comparison of validation methods from a study using 300 simulated datasets and a real dataset of 95 amine mutagens, with models built using LASSO regression [80].
| Validation Technique | Stability (Variation) | Recommended Use Case | Key Advantages | Key Limitations |
|---|---|---|---|---|
| Leave-One-Out (LOO) Cross-Validation | Low variation | High-dimensional data with a small sample size (n << p) [80] | Makes efficient use of limited data; provides a stable performance estimate [80] | Computationally expensive for very large datasets |
| K-Fold Cross-Validation | Moderate variation | Medium to large datasets; general model tuning and validation [84] | Good balance of bias and variance; less computationally intensive than LOO | Performance estimate can have higher variance than LOO for small n [80] |
| External Validation (Single Split) | High variation [80] | Final evaluation of a completely finalized model with sufficient data [3] | Simulates a real-world scenario of predicting truly new compounds | Unreliable for small datasets due to high dependence on a single data split [80] |
| Multi-Split Validation | Moderate to Low variation | Providing a more robust assessment of external predictive ability [80] | Reduces the bias and instability of a single split by averaging over multiple splits | More computationally intensive than a single split |
Experimental Protocol: Comparing Validation Techniques
This protocol outlines a methodology for a comparative study of validation techniques in QSAR modeling, based on published research [80] [83].
1. Dataset Curation and Preparation
- Data Source: Compile a dataset of chemical structures and their associated biological activities from a reliable chemogenomics database like ChEMBL [83].
- Data Cleaning: Standardize chemical structures (e.g., remove salts, neutralize charges, handle tautomers) using tools like the ChemAxon standardizer or RDKit [83].
- Descriptor Calculation: Calculate a large set of molecular descriptors (e.g., ECFP_4 fingerprints or Dragon descriptors) for each compound to create a high-dimensional data matrix [3] [83].
2. Model Building with LASSO Regression
- Algorithm: Use LASSO (Least Absolute Shrinkage and Selection Operator) regression to build the QSAR models. LASSO is suitable for this comparison because it performs automatic variable selection by shrinking the coefficients of irrelevant descriptors to zero, which helps prevent overfitting [80].
- Model Training: Train the LASSO model on the training set determined by each validation method.
3. Application of Validation Techniques Apply the following validation methods to the same dataset and model to allow for a direct comparison:
- Leave-One-Out (LOO) CV: Iteratively use each compound as a test set, training the model on all remaining compounds. The performance metric is averaged over all iterations [80] [84].
- K-Fold CV: Randomly split the dataset into k subsets (folds). For each fold, train the model on the other k-1 folds and use the held-out fold for validation. The reported performance is the average over the k folds [84].
- External Validation: Perform a single random split of the data into a training set (e.g., 80%) and an external test set (e.g., 20%). Train the model on the training set and calculate performance metrics exclusively on the test set [3].
- Multi-Split Validation: Repeat the external validation process multiple times (e.g., 50-100 times) with different random splits. Report the average and variation of the performance metrics [80].
4. Performance Evaluation and Comparison
- For each validation method, calculate predictive performance metrics such as the coefficient of determination (R²) or Root Mean Square Error (RMSE).
- Compare the stability (variation of the metrics) and the average performance across the different methods. The study by Majumdar et al. suggests that for small-sample data, LOO provides the most stable performance estimate [80].
Workflow Diagram for Validation Comparison
The diagram below illustrates the logical workflow for comparing the different validation techniques.
The Scientist's Toolkit: Essential Reagents & Software
The table below lists key software tools and resources essential for conducting QSAR modeling and validation studies.
| Tool/Resource | Function in QSAR Validation | Application Context |
|---|---|---|
| RDKit | Open-source cheminformatics library; used for calculating molecular descriptors (e.g., fingerprints), standardizing structures, and integrating with ML workflows [83] | Descriptor calculation and data preprocessing |
| scikit-learn | A core Python library for machine learning; provides implementations for LASSO, k-fold CV, LOO CV, and train-test splitting [84] | Model building and applying validation techniques |
| Dragon / PaDEL | Software dedicated to calculating thousands of molecular descriptors from chemical structures [3] | High-throughput descriptor calculation |
| ChEMBL Database | A large-scale bioactivity database; provides curated data for building training and test sets for QSAR models [83] | Dataset curation and compilation |
| CORAL Software | A free online tool for building QSAR models using SMILES-based descriptors; useful for robust, validated model development [85] | An alternative approach to QSAR modeling and validation |
Frequently Asked Questions: A QSAR Modeler's Troubleshooting Guide
Model Interpretation & Mechanistic Insight
Q: My QSAR model has good statistical performance, but I cannot explain why the key descriptors are relevant to the biological endpoint. How can I improve mechanistic interpretability?
A: This is a common challenge, especially with complex "black-box" models. The OECD principles emphasize that "a mechanistic interpretation, if possible" is desirable for model acceptance [25] [86]. To address this:
- Start Simple: Before using complex non-linear models, establish a baseline with interpretable linear models like Multiple Linear Regression (MLR) or Partial Least Squares (PLS). These provide clear coefficient estimates for each descriptor, showing the direction and magnitude of their effect on the activity [3].
- Incorporate Domain Knowledge: Relate the selected molecular descriptors to known biological mechanisms or structural alerts. For example, if modeling carcinogenicity, a descriptor like
nRNNOx(number of N-nitroso groups) can be linked to the known structural alert "alkyl and aryl–N-nitroso groups" which can form DNA adducts after metabolic activation [25]. - Use Advanced Interpretable ML: Consider methods specifically designed for interpretability. Recent research has modified Counter-propagation Artificial Neural Networks (CPANN) to dynamically adjust molecular descriptor importance during training, helping to identify which structural features are most critical for classifying specific endpoint classes [25].
Q: What are the most common pitfalls in descriptor selection that hinder mechanistic interpretation?
A: Several pitfalls can obscure the mechanistic meaning of your model:
- Using Too Many Descriptors: Generating hundreds or thousands of descriptors without proper feature selection leads to models that are difficult to interpret and prone to overfitting. Always use feature selection techniques (filter, wrapper, or embedded methods) to identify the most relevant subset [3] [87].
- Ignoring Collinearity: Highly correlated descriptors can produce statistically unstable models where the importance of individual variables is masked. Techniques like Partial Least Squares (PLS) can handle multicollinearity [3].
- Lacking Chemical Intuition: Selecting descriptors based solely on statistical metrics without considering their potential chemical or biological meaning. Always ask: "Does it make sense that this molecular property influences the activity in this way?" [87].
Applicability Domain Assessment
Q: How can I define the Applicability Domain (AD) for my QSAR model to ensure reliable predictions?
A: The Applicability Domain defines the chemical space within which the model can make reliable predictions [86]. It is a critical principle for regulatory acceptance [86]. You can define it using:
- Descriptor Ranges: The most straightforward method is to define the bounds of each descriptor in the training set. A new compound is within the AD if all its descriptor values fall within these ranges [3].
- Leverage and Distance Measures: Calculate the leverage of a new compound relative to the training set. A high leverage indicates the compound is structurally distant from the training data and its prediction may be unreliable [3].
- Consensus Approaches: Use multiple methods (e.g., ranges, PCA, clustering) to get a more robust assessment of the AD. A clear AD definition is essential for determining when your model's predictions should be trusted [3] [86].
Q: What should I do when I need to predict a compound that falls outside my model's Applicability Domain?
A: If a compound falls outside the AD, the prediction should be treated as unreliable [86]. Your options are:
- Do Not Use the Prediction: The safest course of action is to disregard the model's output for this compound.
- Expand Your Training Set (with caution): If feasible and scientifically justified, incorporate more compounds that represent the new chemical space, then rebuild and re-validate the model.
- Use an Alternative Model: Seek or develop a different QSAR model whose Applicability Domain includes your compound of interest. The OECD (Q)SAR Toolbox can be helpful for this [88].
Data Quality & Preparation
Q: My dataset is compiled from multiple sources with varying experimental conditions. How does this affect my QSAR model, and how can I mitigate the issues?
A: Inconsistent biological data is a major pitfall that can lead to models with poor predictive power [87]. The biological data used in a QSAR "should be of a known (and preferably high) quality" [87].
Mitigation Strategies:
- Standardize Biological Activities: Convert all activities to a common unit and scale (e.g., pIC50, pKi) to ensure comparability [3].
- Curate Chemical Structures: Standardize structures by removing salts, normalizing tautomers, and handling stereochemistry consistently [3].
- Document Experimental Conditions: Keep detailed records of the data sources and experimental conditions. If certain data points come from significantly different protocols, consider separating them or applying careful weighting [3] [87].
Model Validation & Regulatory Acceptance
Q: What is the minimum validation required for a QSAR model to be considered reliable for regulatory assessment?
A: The OECD principles require "appropriate measures of goodness-of-fit, robustness, and predictivity" [86]. A robust validation framework includes:
- Internal Validation: Use k-fold cross-validation (e.g., 10-fold) or leave-one-out (LOO) CV on your training set to assess model robustness [3] [86].
- External Validation: This is the gold standard. The model must be tested on a fully independent external test set of compounds that were not used in any part of the model development process. This provides a realistic estimate of its predictive power on new data [3] [87] [86].
- Statistical Metrics: Report multiple metrics such as R² (coefficient of determination), RMSE (root mean square error), and AIC (Akaike Information Criterion) for both the training and test sets to give a comprehensive view of model performance [86].
Experimental Protocols for Robust QSAR
Protocol 1: Building an OECD-Compliant QSAR Model
This protocol provides a step-by-step methodology for developing a QSAR model that aligns with OECD principles [3] [86].
Dataset Curation & Preparation
- Source: Collect chemical structures and associated biological activities from reliable, well-documented sources (e.g., ChEMBL, ZINC) [86].
- Clean: Remove duplicates and errors. Standardize structures (e.g., neutralize charges, remove salts) [3].
- Scale: Convert biological activities to a consistent scale (e.g., pKi). Scale molecular descriptors to have zero mean and unit variance [3].
- Split: Divide the data into a training set (for model building), an optional validation set (for hyperparameter tuning), and a hold-out external test set (for final evaluation). The test set must remain completely unused until the final model is built [3] [86].
Molecular Descriptor Calculation & Selection
- Calculate: Use software like PaDEL-Descriptor, RDKit, or Mordred to compute a wide range of 2D and/or 3D molecular descriptors [3] [86].
- Select: Apply feature selection techniques (e.g., genetic algorithms, LASSO regression, correlation analysis) to reduce the number of descriptors and minimize overfitting. The goal is a parsimonious set of mechanistically interpretable descriptors [3] [86].
Model Building & Training
- Algorithm Selection: Choose an algorithm based on your data and need for interpretability. Start with MLR or PLS for simplicity and interpretability. Progress to Support Vector Machines (SVM) or Neural Networks if non-linearity is suspected and data volume is sufficient [3].
- Train: Build the model using only the training set.
- Internal Validation: Perform 10-fold cross-validation on the training set to tune parameters and estimate robustness [86].
Model Validation & Documentation
- External Test: Apply the final model to the held-out external test set to obtain the best estimate of its predictive ability [86].
- Define Applicability Domain: Clearly define the chemical space of the model using descriptor ranges or other distance measures [86].
- Interpret Mechanistically: Analyze the model to explain how the key descriptors influence the activity, linking them to known mechanisms or physicochemical principles where possible [25] [86].
Protocol 2: Assessing Mechanistic Interpretability using CPANN
This protocol is based on a recent study that enhanced the interpretability of neural network models for classification endpoints [25].
- Dataset: Obtain a curated dataset with a classification endpoint (e.g., hepatotoxicity, enzyme inhibition). The hepatotoxicity dataset from the LiverTox database, described using 49 molecular descriptors, is an example [25].
- Model Setup: Configure a Counter-propagation Artificial Neural Network (CPANN). The architecture consists of two layers: a Kohonen layer (for grouping molecules by structural similarity) and a Grossberg layer (for predicting the target property) [25].
- Training with Dynamic Importance: Implement the modified CPANN algorithm that dynamically adjusts the importance (
m(t, i, j, k)) of each molecular descriptor (k) on each neuron during training. This adjustment is based on the difference between the input object's descriptor/target values and the neuron's weights [25]. - Interpretation: After training, analyze the network to identify which descriptors were most frequently assigned high importance for classifying molecules into specific endpoint classes. This can reveal sub-structural features or properties critical for the activity [25].
QSAR Model Development Workflow
The following diagram illustrates the integrated, iterative workflow for developing a QSAR model that fulfills the OECD principles, highlighting how mechanistic interpretation and applicability domain assessment inform the process.
The Scientist's Toolkit: Essential Research Reagents & Software
The following table details key software tools and conceptual "reagents" essential for conducting QSAR research in line with OECD principles.
| Tool / Reagent | Primary Function | Relevance to OECD Principles & Descriptor Selection |
|---|---|---|
| PaDEL-Descriptor [3] | Calculates molecular descriptors and fingerprints. | Generates a wide array of constitutional, topological, and electronic descriptors for building a initial descriptor pool. |
| RDKit / Mordred [3] [86] | Open-source cheminformatics toolkits for descriptor calculation. | Provides computational "reagents" to numerically encode molecular structures, forming the basis of the QSAR model. |
| OECD (Q)SAR Toolbox [88] | A software application that helps to fill data gaps by grouping chemicals into categories. | Aids in assessing the chemical space and read-across, supporting the definition of the Applicability Domain. |
| AutoML Platforms (e.g., H2O) [86] | Automates the process of algorithm selection, feature engineering, and model tuning. | Helps achieve "an unambiguous algorithm" and "appropriate measures of... predictivity" by systematically optimizing the model building process. |
| Interpretable ML Methods (e.g., CPANN-v2) [25] | Neural network algorithms designed to provide insight into descriptor importance. | Directly addresses the principle of "a mechanistic interpretation" by identifying which structural features drive classification. |
Frequently Asked Questions (FAQs)
FAQ 1: What are the key performance differences between 2D, 3D, and combined descriptor sets in QSAR modeling?
Multiple studies have demonstrated that combining different descriptor types typically yields superior performance. A 2023 comparison based on bioactive conformations found that while 2D and 3D descriptors individually produced significant models, combining them resulted in "many more significant models" due to their ability to encode "different, yet complementary molecular properties" [89]. Similarly, a 2022 benchmark on ADME-Tox targets showed that traditional 1D, 2D, and 3D descriptors generally outperformed fingerprint-based methods when used with the XGBoost algorithm [90].
Table 1: Performance Comparison of Descriptor Types Across Studies
| Descriptor Type | Key Strengths | Performance Notes | Ideal Use Cases |
|---|---|---|---|
| 2D Descriptors | Fast calculation; No conformation needed; Good for scaffold hopping | Often performs nearly as well as combined sets [90] | Preliminary screening; Large virtual libraries |
| 3D Descriptors | Encodes spatial information; Captures stereochemistry | Performance gains when bio-active conformation is known [89] | Target-specific modeling; Protein-ligand interaction |
| Descriptor Combinations (2D+3D) | Complementary information; More comprehensive representation | "Many more significant models" than single-type descriptors [89] | Lead optimization; High-precision prediction |
| AI-Generated Descriptors | Data-driven features; No manual engineering | Captures abstract hierarchical features [41] | Complex endpoint prediction; Large diverse datasets |
FAQ 2: How do AI-generated descriptors compare to traditional molecular descriptors?
AI-generated "deep descriptors" represent a paradigm shift from manually engineered features to learned representations. According to recent reviews, graph neural networks (GNNs) and other deep learning approaches create "latent embeddings" that capture "more abstract and hierarchical molecular features" without manual descriptor engineering [41]. These data-driven descriptors are particularly valuable for complex endpoints where the relevant structural features are not fully understood, enabling the construction of "flexible QSAR pipelines applicable across diverse chemical spaces" [41]. However, these models often face challenges in interpretability compared to traditional descriptors [91].
FAQ 3: What is the impact of descriptor preselection and intercorrelation limits on model performance?
Descriptor preselection is a critical step that significantly impacts model quality and stability. A 2019 systematic study examined this effect across four case studies and found that the choice of intercorrelation limit (the threshold for removing highly correlated descriptors) is dataset-dependent [92]. The research concluded that while there's no universal optimal value, applying some rational intercorrelation limit (commonly between 0.90-0.95) generally improves model robustness compared to either no filtering or extremely strict limits [92]. The removal of constant or near-constant descriptors is also standard practice to reduce noise in the feature set [90].
Troubleshooting Guides
Issue 1: Poor Model Generalization Despite High Training Accuracy
Problem: Your QSAR model performs well on training data but poorly on external test sets or new chemical domains.
Solution: Implement rigorous benchmark validation using synthetic datasets with known ground truth.
Utilize Predefined Pattern Benchmarks: Employ benchmark datasets where endpoints are determined by predefined patterns, enabling you to verify if your model can retrieve these known patterns [91]. Examples include:
- Simple additive properties: Where specific contributions are assigned to individual atoms.
- Context-dependent properties: Where contributions depend on local chemical environments.
- Pharmacophore-like settings: Where activity depends on specific 3D patterns [91].
Apply Quantitative Interpretation Metrics: Use metrics like SRD (Sum of Ranking Differences) combined with ANOVA to quantitatively compare model performance and identify the most robust descriptor sets for your specific data [92].
Check Descriptor Intercorrelation: Apply an intercorrelation limit (e.g., 0.90-0.95) to remove redundant descriptors, which can improve model stability and generalizability [92].
Table 2: Essential Research Reagents & Computational Tools
| Tool / Resource | Type | Primary Function | Access |
|---|---|---|---|
| DRAGON | Software | Calculates >4000 molecular descriptors (1D-3D) | Commercial |
| QSARINS | Software | MLR modeling with genetic algorithm variable selection | Open Access |
| RDKit | Cheminformatics Library | Fingerprint generation (Morgan), descriptor calculation | Open Source |
| Benchmark Datasets [91] | Data | Synthetic datasets with pre-defined structure-activity rules | Open Access |
| MedMNIST v2 [93] | Data | Standardized 2D/3D biomedical image classification benchmark | Open Access |
Issue 2: Inconsistent Performance Across Different Chemical Domains or Target Classes
Problem: Your descriptor set works well for one target or chemical series but fails to maintain performance across diverse datasets.
Solution: Adopt a hybrid descriptor strategy and evaluate across multiple benchmark endpoints.
Combine Descriptor Types: Integrate 2D and 3D descriptors to capture complementary information, as studies consistently show combined approaches outperform single-type descriptors [89] [90].
Benchmark Across Diverse Tasks: Test descriptor performance across multiple ADME-Tox targets (e.g., Ames mutagenicity, hERG inhibition, BBB permeability) to identify universally robust descriptors or context-specific strengths [90].
Evaluate AI Descriptors for Complex Problems: For particularly challenging endpoints with complex structure-activity relationships, implement graph neural networks or transformer-based models that generate task-optimized descriptors rather than relying on pre-defined features [41].
Troubleshooting Workflow for Descriptor Performance Issues
Issue 3: Difficulty Interpreting Complex "Black Box" Models, Especially with AI-Generated Descriptors
Problem: Your models produce accurate predictions but offer little insight into the structural features driving activity, making it difficult to guide chemical optimization.
Solution: Implement modern interpretation frameworks specifically designed for complex models.
Leverage Model-Agnostic Interpretation Tools: Apply methods like SHAP (SHapley Additive exPlanations) and LIME (Local Interpretable Model-agnostic Explanations) that can explain predictions from any model, regardless of descriptor type [41].
Use Structural Interpretation Benchmarks: Validate your interpretation methods on benchmark datasets with known structure-activity relationships (e.g., where activity depends on specific functional groups) to ensure they correctly identify contributing motifs [91].
Focus on Explainable AI Approaches: When using deep learning models, prioritize architectures with built-in interpretability, such as attention mechanisms that can highlight important atoms or fragments directly from molecular structures [41] [91].
Frequently Asked Questions
FAQ: What are the most critical steps in QSAR model development to ensure regulatory acceptance for REACH?
The most critical steps involve rigorous validation and clearly defining the applicability domain of your model. For REACH compliance, the European Chemicals Agency (ECHA) requires QSAR models to be scientifically robust and reliable. This is achieved through internal validation (e.g., cross-validation) and external validation using a separate test set of compounds. Furthermore, you must clearly define the chemical space for which your model makes reliable predictions. The leverage method is a common way to define this domain and identify when you are predicting compounds that are too structurally dissimilar from your training data [43].
FAQ: How does descriptor intercorrelation (multicollinearity) affect my QSAR model, and how can I address it?
Descriptor intercorrelation occurs when two or more predictor variables are highly correlated, making it difficult to determine their individual effects on the biological activity. This can lead to overfitting and models that perform poorly on new, unseen data [72]. To address this, you can:
- Use algorithms robust to multicollinearity, such as Gradient Boosting models, which naturally prioritize informative descriptors and down-weight redundant ones [72].
- Apply feature selection methods like Recursive Feature Elimination (RFE), which iteratively removes the least important descriptors [41] [72].
- Calculate a correlation matrix for your descriptors and remove those with very high correlation to others, though this simple method may sometimes discard useful information [72].
FAQ: What is the difference between classical and machine learning QSAR models in terms of interpretability for regulatory submissions?
Classical QSAR models, such as those built with Multiple Linear Regression (MLR), are often preferred in regulatory settings like REACH because of their simplicity and ease of explanation. The relationship between descriptors and activity is transparent in a linear equation, which aids in mechanistic interpretation and compliance [41] [43]. In contrast, complex machine learning models like Artificial Neural Networks (ANNs) can be "black boxes," making it harder to explain which structural features drive the prediction. However, methods like SHAP (SHapley Additive exPlanations) are increasingly used to interpret these complex models and provide the necessary transparency for regulatory acceptance [41] [25].
FAQ: My QSAR model passed validation but failed to predict a new compound accurately. What might have gone wrong?
This is a classic sign that the new compound falls outside the applicability domain of your model. Even a well-validated model is only reliable for predicting compounds that are structurally similar to those it was trained on. The new compound may possess functional groups, descriptor values, or structural features not represented in your original training set. Always use the applicability domain to screen new compounds before running predictions [43] [94].
Troubleshooting Guides
Issue: Model is Overfitting the Training Data
Problem: Your QSAR model shows excellent performance on the training data but performs poorly on the validation or test set.
Solutions:
- Simplify the Model: Reduce the number of molecular descriptors. Use feature selection techniques like RFE or genetic algorithms to identify the most predictive subset of descriptors [41] [72] [25].
- Use Robust Algorithms: Switch to machine learning methods known for handling overfitting, such as Random Forests or Gradient Boosting. These ensemble methods are less prone to overfitting than simpler models when dealing with high-dimensional data [41] [72].
- Increase Training Data: If possible, add more diverse compounds to your training set to better represent the chemical space.
- Apply Regularization: Use techniques like LASSO (Least Absolute Shrinkage and Selection Operator), which penalizes model complexity and can automatically drive less important descriptor coefficients to zero [41].
Issue: Poor Predictive Power (Underfitting)
Problem: The model performs poorly on both training and test data, indicating it failed to learn the underlying structure-activity relationship.
Solutions:
- Check Feature Selection: You may have removed too many descriptors or kept irrelevant ones. Re-evaluate your descriptor set using mutual information or other importance rankings [41].
- Explore Non-Linear Models: The relationship between your descriptors and the activity may be non-linear. Try using Artificial Neural Networks (ANNs), Support Vector Machines (SVM) with non-linear kernels, or Gradient Boosting, which can capture these complex patterns [43] [72].
- Engineer New Descriptors: The current descriptor set may not capture the relevant chemical information. Consider calculating additional descriptors, such as 3D field descriptors (e.g., Cresset XED) or quantum chemical descriptors (e.g., HOMO-LUMO energy gap) [41] [72].
Issue: Model Fails REACH Compliance Check
Problem: A QSAR model submitted for REACH compliance is rejected due to insufficient validation or lack of mechanistic interpretability.
Solutions:
- Follow OECD Principles: Ensure your model adheres to the five OECD principles for QSAR validation, which include a defined endpoint, an unambiguous algorithm, a defined domain of applicability, appropriate measures of goodness-of-fit and robustness, and a mechanistic interpretation, if possible [25].
- Document the Applicability Domain: Explicitly define and document the applicability domain using methods like the leverage approach. Provide a clear description of the chemical space covered by your training set [43].
- Enhance Interpretability: Even for complex models, use post-hoc interpretation tools like SHAP or LIME to identify which molecular features contribute most to the prediction. This can provide the "mechanistic interpretation" encouraged by OECD guidelines [41] [25].
- Provide Comprehensive Validation Metrics: Do not rely solely on R². Include a full suite of validation metrics from both internal and external validation, such as Q², RMSE, and MAE, for both training and test sets [43].
Experimental Protocols & Methodologies
Protocol 1: Developing a Validated MLR QSAR Model for REACH
This protocol outlines the steps for building a interpretable Multiple Linear Regression (MLR) QSAR model, suitable for regulatory submissions.
1. Data Curation and Preparation
- Collect a homogeneous set of compounds with experimentally measured biological activity (e.g., IC50) obtained from a standardized assay [43].
- Divide the dataset randomly into a training set (~70-80%) for model development and a test set (~20-30%) for external validation. A common practice is to use about 2/3 of the data for training [43].
2. Molecular Descriptor Calculation and Pre-processing
- Calculate a wide range of molecular descriptors (e.g., physicochemical, topological) using software like RDKit, PaDEL, or DRAGON [41].
- Pre-process the descriptors: Remove those with constant or near-constant values and those with a high number of missing values. Impute missing values if appropriate and robust to do so.
3. Descriptor Selection and Model Training
- Use stepwise regression or genetic algorithms to select a subset of descriptors that are statistically significant and not highly correlated with each other [41] [43].
- Construct the MLR model using the training set data, resulting in a simple linear equation:
Activity = C + (a × D1) + (b × D2) + ...[43].
4. Model Validation and Defining Applicability Domain
- Internal Validation: Perform leave-one-out (LOO) or leave-many-out (LMO) cross-validation on the training set to calculate Q², which assesses model robustness [43].
- External Validation: Use the held-out test set to evaluate the model's predictive power. Calculate predictive R² and RMSE for the test set [43].
- Define Applicability Domain: Use the leverage method. Calculate the leverage (h) for each compound. The critical leverage value (h) is typically set to 3p/n, where p is the number of model descriptors and n is the number of training compounds. A new compound with a leverage higher than h is considered outside the applicability domain [43].
Protocol 2: Building a Non-Linear ANN QSAR Model with Enhanced Interpretation
This protocol is for developing a high-performance, non-linear model using Artificial Neural Networks (ANNs) and then interpreting it for regulatory contexts.
1. Data Curation and Preparation
- Follow the same steps as Protocol 1 for data collection and splitting into training and test sets.
2. Molecular Descriptor Calculation and Pre-processing
- Calculate a large pool of molecular descriptors.
- Pre-process the data by scaling descriptors (e.g., standard scaling) to a common range, which is critical for the performance of ANNs and many other machine learning algorithms [72].
3. Model Training with Hyperparameter Optimization
- Design an ANN architecture (e.g., [8.11.11.1] denoting 8 input descriptors, two hidden layers with 11 neurons each, and 1 output neuron) [43].
- Use a hold-out validation set from the training data or cross-validation to tune hyperparameters (e.g., learning rate, number of layers and neurons, epochs) to prevent overfitting and find the optimal model.
4. Model Interpretation and Validation
- Validation: Conduct rigorous internal and external validation as described in Protocol 1.
- Interpretation: Apply model interpretation techniques like SHAP to determine the contribution of each molecular descriptor to the final predicted activity. This helps transform the "black box" model into an interpretable tool, identifying key structural features that drive activity [41] [25].
Table 1: Key Validation Metrics for QSAR Models
| Metric | Formula / Description | Acceptance Threshold Guideline | Purpose |
|---|---|---|---|
| R² (Coefficient of Determination) | R² = 1 - (SS₍res₎/SS₍tot₎) | > 0.6 | Measures goodness-of-fit of the model to the training data [43] |
| Q² (Cross-validated R²) | Q² = 1 - (PRESS/SS₍tot₎) | > 0.5 | Assesses internal robustness and predictive ability within the training set via cross-validation [43] |
| RMSE (Root Mean Square Error) | RMSE = √(Σ(Ŷᵢ - Yᵢ)²/n) | As low as possible | Measures the average difference between predicted and experimental values; lower is better [72] |
| Applicability Domain (Leverage) | h* = 3p/n | New compound with h < h* | Defines the chemical space where the model is reliable; h is the leverage of a new compound [43] |
Table 2: The Scientist's Toolkit: Essential Research Reagents & Software for QSAR
| Tool / Reagent | Type | Primary Function in QSAR |
|---|---|---|
| RDKit | Software/Cheminformatics Library | Open-source toolkit for cheminformatics; used for calculating 2D molecular descriptors and fingerprints [41] [72] |
| IUCLID | Software/Regulatory Tool | Software required for submitting registration dossiers to ECHA under REACH [95] [96] |
| SHAP (SHapley Additive exPlanations) | Software/Interpretation Library | A game-theoretic method to explain the output of any machine learning model, providing descriptor importance for individual predictions [41] [25] |
| Curated Experimental Bioactivity Datasets | Data/Reagent | High-quality, standardized biological data (e.g., from public databases like LiverTox) used as the foundation for training and testing QSAR models [25] |
| Cresset XED Field Descriptors | Software/Computational Descriptor | 3D molecular descriptors that model a ligand's shape and electrostatic character as a protein would "see" it, used in 3D-QSAR [72] |
Experimental Workflow Diagrams
Descriptor Selection and Validation Workflow for Regulatory QSAR
Integrating QSAR into the REACH Compliance Process
Conclusion
The strategic selection of molecular descriptors is the cornerstone of developing robust, predictive, and interpretable QSAR models. This synthesis underscores that no single descriptor type is universally superior; the optimal choice is dictated by the specific endpoint, chemical space, and desired balance between interpretability and predictive accuracy. The future of descriptor selection lies in the intelligent integration of AI and machine learning, which can dynamically adjust descriptor importance and generate insightful latent representations. As the field advances with larger, higher-quality datasets and more sophisticated algorithms, a principled approach to descriptor selection—grounded in rigorous validation and a clear understanding of the model's applicability domain—will be paramount. This will accelerate the discovery of novel therapeutics and enhance the reliability of environmental risk assessments, solidifying QSAR's vital role in biomedical and clinical research.
References
- 1. Molecular descriptor [https://en.wikipedia.org/wiki/Molecular_descriptor]
- 2. Computing Molecular Descriptors - Part 1 [https://drzinph.com/computing-molecular-descriptors-intro/]
- 3. A Beginner's Guide to QSAR Modeling in Cheminformatics ... [https://neovarsity.org/blogs/cheminformatics-qsar-beginner-guides]
- 4. Quantitative structure-activity relationship (QSAR) studies ... [https://www.sciencedirect.com/science/article/pii/S3050787125000186]
- 5. Causal Descriptors in QSAR: Deconfounding High ... [https://chemrxiv.org/engage/chemrxiv/article-details/68fffd90113cc7cfff76df91]
- 6. Hierarchic system of QSAR models (1D-4D) on the base ... [https://pubmed.ncbi.nlm.nih.gov/16237516/]
- 7. Utilizing Molecular Descriptor Importance to Enhance ... [https://pubmed.ncbi.nlm.nih.gov/40423462/]
- 8. QSAR Toolbox [https://qsartoolbox.org/support/]
- 9. FAQ - QSAR Lab [https://www.qsarlab.com/en/faq/]
- 10. IN HANSCH in principles of drug discovery | PPTX ANALYSIS QSAR [https://www.slideshare.net/slideshow/hansch-analysis-in-qsar-in-principles-of-drug-discovery/283929012]
- 11. Experimental Errors in QSAR Modeling Sets: What We Can ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5494643/]
- 12. Molecular similarity in chemical informatics and predictive ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12010357/]
- 13. Sciome to Spotlight Breakthroughs in Read-Across and ... [https://www.sciome.com/sciome-to-spotlight-breakthroughs-in-read-across-and-qsar-at-ascct-2025/]
- 14. Revisiting the Use of Quantum Chemical Calculations in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9866719/]
- 15. Lipophilicity - an overview [https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/lipophilicity]
- 16. Quantitative study of electrostatic and steric effects on ... [https://www.sciencedirect.com/science/article/abs/pii/S1093326305000720]
- 17. Tutorial: Molecular Descriptors in QSAR [https://people.chem.ucsb.edu/kahn/kalju/chem162/public/molecules_qsar.html]
- 18. Steric parameters, molecular modeling and hydropathic ... [https://pubmed.ncbi.nlm.nih.gov/25522019/]
- 19. QSAR analysis of salicylamide isosteres with the use of ... [https://pubmed.ncbi.nlm.nih.gov/18547684/]
- 20. Application of topological descriptors in QSAR and drug ... [https://pubmed.ncbi.nlm.nih.gov/12462157/]
- 21. Descriptors and their selection methods in QSAR analysis [https://www.sciencedirect.com/science/article/abs/pii/S1359644616302318]
- 22. Quantum chemical descriptors in quantitative structure ... [https://www.sciencedirect.com/science/article/abs/pii/S0169743921001520]
- 23. The Internal Relation between Quantum Chemical Descriptors ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6278375/]
- 24. Recent Developments in 3D QSAR and Molecular Docking ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7123761/]
- 25. Utilizing Molecular Descriptor Importance to Enhance ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12115611/]
- 26. QSAR-guided discovery of novel KRAS inhibitors for lung ... [https://www.frontiersin.org/journals/bioinformatics/articles/10.3389/fbinf.2025.1663846/full]
- 27. A review of quantitative structure-activity relationship [https://www.sciencedirect.com/science/article/abs/pii/S0169743924002181]
- 28. Application of the LAD-LASSO as a dimensional reduction ... [https://www.sciencedirect.com/science/article/abs/pii/S0169743922000211]
- 29. The Role of Mutual Information Estimator Choice in Feature ... [https://www.mdpi.com/2078-2489/16/9/724]
- 30. Feature selection using Decomposed Mutual Information ... [https://www.sciencedirect.com/science/article/pii/S0925231222011936]
- 31. Comparison of commercially available genetic algorithms [https://link.springer.com/article/10.1007/s10822-004-5322-1]
- 32. Comparison of various methods for validity evaluation of ... [https://bmcchem.biomedcentral.com/articles/10.1186/s13065-022-00856-4]
- 33. Variable Selection via Fused Sparse‐Group Lasso ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12559784/]
- 34. QSAR Models for Predicting the Antioxidant Potential of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12194667/]
- 35. Dimensionality Reduction and Deep Dive Into Principal ... [https://medium.com/data-science/deep-dive-into-principal-component-analysis-fc64347c4d20]
- 36. Principal Component Analysis for Dimensionality ... [https://www.machinelearningmastery.com/principal-components-analysis-for-dimensionality-reduction-in-python/]
- 37. Practical Example of Dimensionality Reduction [https://towardsdatascience.com/practical-example-of-dimensionality-reduction-d0525632c355/]
- 38. Principal Component Analysis (PCA) of Molecular ... [https://www.mdpi.com/1422-0067/25/1/192]
- 39. Design, Quantitative Structure–Activity Relationships and ... [https://www.sciencedirect.com/science/article/pii/S2667022425001458]
- 40. GCAC: galaxy workflow system for predictive model building ... [https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-018-2492-8]
- 41. AI-Integrated QSAR Modeling for Enhanced Drug Discovery [https://pmc.ncbi.nlm.nih.gov/articles/PMC12525248/]
- 42. NfκBin: a machine learning based method for screening TNF ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12310657/]
- 43. Advancing QSAR models in drug discovery for best ... [https://www.sciencedirect.com/science/article/abs/pii/S0169743925002291]
- 44. QSAR Modeling of Histamine H3R Antagonists/inverse ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6080103/]
- 45. Comparative Study of (Q)SAR Models for Predicting the ... [https://www.sciencedirect.com/science/article/pii/S3050620425000491]
- 46. When to Use QSAR for Chemical Substance Registration ... [https://www.chemsafetypro.com/Topics/CRA/When_to_Use_QSAR_for_Chemical_Substance_Registration_and_Common_QSAR_Models.html]
- 47. VEGA interpretation [https://www.vegahub.eu/download/vega-interpretation/]
- 48. EPI Suite™-Estimation Program Interface [https://www.epa.gov/tsca-screening-tools/epi-suitetm-estimation-program-interface]
- 49. Interpretation-ADMElab: ADMET Prediction [https://admet.scbdd.com/home/interpretation/]
- 50. Comprehensive benchmarking of computational tools for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11674477/]
- 51. Representative feature selection of molecular descriptors ... [https://www.sciencedirect.com/science/article/abs/pii/S0022286021013788]
- 52. Bio-QSARs 2.0: Unlocking a new level of predictive power ... [https://www.sciencedirect.com/science/article/pii/S0160412024001934]
- 53. Topological regression as an interpretable and efficient ... [https://www.nature.com/articles/s41467-024-49372-0]
- 54. A Review of Quantitative Structure-Activity Relationship ... [https://pubmed.ncbi.nlm.nih.gov/41012420/]
- 55. A (Comprehensive) Review of the Application ... [https://www.mdpi.com/2076-3417/15/3/1206]
- 56. Duality of Simplicity and Accuracy in QSPR [https://www.mdpi.com/1420-3049/30/22/4361]
- 57. 2D and 3D QSAR-Based Machine Learning Models for ... [https://www.sciencedirect.com/science/article/pii/S2468227625005381]
- 58. One size does not fit all: revising traditional paradigms for ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-00948-y]
- 59. Extrapolative prediction of small-data molecular property ... [https://www.nature.com/articles/s41524-023-01194-2]
- 60. Improving the accuracy of prediction models for small datasets ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-01015-2]
- 61. Insight into Machine Learning Models to Predict Toxicity of ... [https://www.sciencedirect.com/science/article/pii/S2405665025001337]
- 62. For drug discovery, QSAR and molecular descriptors are ... [https://www.linkedin.com/pulse/drug-discovery-qsar-molecular-descriptors-more-relevant-vijay-masand-vdycf]
- 63. The Generation and Validation of QSAR Model [https://bio-protocol.org/exchange/minidetail?id=9718697&type=30]
- 64. Unified and explainable molecular representation learning ... [https://www.nature.com/articles/s41467-025-63730-6]
- 65. Science Brief: 3D-QSAR Machine Learning for Binding ... [https://www.eyesopen.com/science-briefs/3d-qsar-machine-learning]
- 66. A comprehensive in-silico approach (pharmacophoric ... [https://www.sciencedirect.com/science/article/pii/S157018082500017X]
- 67. Comparison of Different Approaches to Define the Applicability Domain of QSAR Models - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6268288/]
- 68. Study of the Applicability Domain of the QSAR Classification Models by Means of the Rivality and Modelability Indexes - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6278359/]
- 69. Prediction reliability of QSAR models: an overview ... [https://pubmed.ncbi.nlm.nih.gov/35267067/]
- 70. Quantitative structure–activity relationship - Wikipedia [https://en.wikipedia.org/wiki/Quantitative_structure%E2%80%93activity_relationship]
- 71. A novel non-linear approach for establishing a QSAR ... [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2023.1263933/full]
- 72. Tackling descriptor intercorrelation in QSAR models | Cresset [https://cresset-group.com/science/science-resources/qsar-models-tackling-descriptor-intercorrelation/]
- 73. An automated framework for QSAR model building [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-017-0256-5]
- 74. AI-Integrated QSAR Modeling for Enhanced Drug Discovery [https://www.mdpi.com/1422-0067/26/19/9384]
- 75. Hybridizing Feature Selection and ... [https://www.nature.com/articles/s41598-017-02114-3]
- 76. Known Issues ⬣ QSAR Toolbox [https://qsartoolbox.org/download/issues/]
- 77. Technical issues - QSAR Toolbox Helpdesk - Freshdesk [https://qsartoolbox.freshdesk.com/support/solutions/folders/16000079662/page/2]
- 78. Explained vs. Predictive Power: R , Adjusted ² , and R ² Beyond [https://www.r-bloggers.com/2025/04/explained-vs-predictive-power-r%C2%B2-adjusted-r%C2%B2-and-beyond/]
- 79. External Model Validation [http://facweb.cs.depaul.edu/sjost/csc423/documents/model-validation.htm]
- 80. Beware of External Validation! - A Comparative Study of Several Validation Techniques used in QSAR Modelling - PubMed [https://pubmed.ncbi.nlm.nih.gov/29701159/]
- 81. Comparison of various methods for validity evaluation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9396839/]
- 82. Descriptors and their selection methods in QSAR analysis: paradigm for drug design - PubMed [https://pubmed.ncbi.nlm.nih.gov/27326911/]
- 83. Utilizing random Forest QSAR models with optimized ... [https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-017-1960-x]
- 84. 3.1. Cross-validation: evaluating estimator performance [https://scikit-learn.org/stable/modules/cross_validation.html]
- 85. validated all-qsar models: Topics by Science.gov [https://www.science.gov/topicpages/v/validated+all-qsar+models]
- 86. Do AutoML-Based QSAR Models Fulfill OECD Principles ... [https://www.mdpi.com/1999-4923/14/7/1415]
- 87. Pitfalls in QSAR [https://www.sciencedirect.com/science/article/abs/pii/S0166128002006164]
- 88. Quantitative Structure-Activity Relationships Project [https://www.oecd.org/en/topics/sub-issues/assessment-of-chemicals/quantitative-structure-activity-relationships-project.html]
- 89. A Comparison between 2D and 3D Descriptors in QSAR ... [https://pubmed.ncbi.nlm.nih.gov/36617991/]
- 90. Comparison of Descriptor- and Fingerprint Sets in Machine ... [https://www.frontiersin.org/journals/chemistry/articles/10.3389/fchem.2022.852893/full]
- 91. Benchmarks for interpretation of QSAR models [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-021-00519-x]
- 92. Intercorrelation Limits in Molecular Descriptor Preselection ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6767540/]
- 93. MedMNIST v2 - A large-scale lightweight benchmark for 2D ... [https://www.nature.com/articles/s41597-022-01721-8]
- 94. What is QSAR and how is it applied in bioinformatics? [https://synapse.patsnap.com/article/what-is-qsar-and-how-is-it-applied-in-bioinformatics]
- 95. EU REACH [https://www.trade.gov/eu-reach]
- 96. REACH Registration: Lead Registrant or Member Registrant? [https://ecomundo.eu/en/blog/understanding-reach-key-points-registration-process]